Smartphone-Based Lab-on-a-Chip: Principles and Advances in Optical Detection for Biomedical Applications
This article provides a comprehensive review of optical detection methods integrated with smartphone-based lab-on-a-chip (LoC) platforms, tailored for researchers and professionals in drug development and biomedical science.
Smartphone-Based Lab-on-a-Chip: Principles and Advances in Optical Detection for Biomedical Applications
Abstract
This article provides a comprehensive review of optical detection methods integrated with smartphone-based lab-on-a-chip (LoC) platforms, tailored for researchers and professionals in drug development and biomedical science. It explores the foundational principles of colorimetric, fluorescence, and label-free optical techniques, detailing their implementation through smartphone cameras and sensors. The scope extends to advanced applications, including single-molecule detection and super-resolution imaging, alongside a critical analysis of real-world challenges such as signal variability, calibration, and system integration. A comparative evaluation of performance metrics, limits of detection, and scalability offers a practical framework for selecting and validating appropriate methods for specific research or diagnostic needs, positioning smartphone-based LoC as a transformative tool for decentralized, point-of-care analysis.
Core Principles and Enabling Technologies of Smartphone Optical Detection
Optical detection methods represent a cornerstone of modern analytical science, particularly within the rapidly evolving field of smartphone-based Lab-on-Chip (LoC) research. These techniques enable the direct, real-time, and label-free detection of biological and chemical substances with high specificity and sensitivity [1]. The integration of optical detection principles with mobile technology has catalyzed a paradigm shift in point-of-care diagnostics, environmental monitoring, and drug development, making sophisticated analytical capabilities accessible in resource-limited settings [2]. This technical guide provides an in-depth examination of three fundamental optical detection methodologies—colorimetric, fluorescence, and interferometric scattering—framed within the context of their implementation in smartphone-based LoC platforms. By elucidating the underlying physics, instrumental configurations, and practical applications of each technique, this review aims to equip researchers and drug development professionals with the knowledge necessary to advance the development of decentralized, mobile-based diagnostic solutions.
Core Principles and Physics of Optical Detection
Optical biosensors function by converting a biological interaction into a quantifiable optical signal, which can be broadly categorized into label-free and label-based detection modalities [1]. In label-free sensing, the detected signal arises directly from the interaction between the analyte and the transducer surface. In contrast, label-based approaches utilize optical tags such as fluorophores or enzymes that generate colorimetric, fluorescent, or luminescent signals upon biological binding events [1]. The dominance of optical detection in biosensing stems from its compatibility with diverse transduction mechanisms, relatively straightforward integration with microfluidic platforms, and capacity for high-sensitivity, multi-analyte detection [3].
The operational principles of optical biosensors often exploit the evanescent field phenomenon, where light propagating through a waveguide generates an electromagnetic field that extends approximately one wavelength into the lower-refractive-index medium surrounding the waveguide [1]. This decaying field is exquisitely sensitive to changes in the interfacial properties, enabling the detection of molecular binding events occurring within this narrow region without interference from bulk solution effects. Surface Plasmon Resonance (SPR), interferometry, and evanescent wave fluorescence all leverage this fundamental principle to achieve exceptional sensitivity for biomolecular interactions [1].
Colorimetric Detection
Fundamental Principle
Colorimetric sensing establishes a quantitative relationship between the concentration of an analyte and specific colorimetric data generated through chromogenic or discoloration reactions [4]. This detection method relies on measurable changes in the absorption of light by a sample, typically quantified using the Beer-Lambert Law, which states that absorbance (A) is proportional to the concentration (c) of the absorbing species and the path length (l) of the light through the sample: A = εlc, where ε is the molar absorptivity coefficient [5]. The color change can be instigated by various mechanisms including enzymatic assays, redox indicators, pH indicators, and nanoparticle aggregation (e.g., gold or silver nanoparticles) [4].
Smartphone Integration and Measurement
Smartphone-based colorimetric detection leverages the device's built-in camera as a spectrometer and its processing capabilities for data analysis [4]. The operational workflow typically involves three key steps:
- Colorimetric Transduction: A biochemical reaction generates a color signal proportional to the analyte concentration.
- Image Capture: The smartphone camera captures an image of the reaction platform (e.g., microtiter plate, paper-based device, microfluidic chip), often using customized accessories to minimize ambient light interference.
- Data Processing: A dedicated application on the smartphone processes the image, typically by converting it to a standard color space such as RGB (Red, Green, Blue), HSV (Hue, Saturation, Value), or CIE LAB*, and fits appropriate mapping relations to quantify the analyte concentration [4].
This approach significantly enhances the portability and accessibility of colorimetric testing, enabling point-of-care and on-site diagnostics outside traditional laboratory settings.
Experimental Protocol: Smartphone-Based Colorimetric Detection
Objective: To quantitatively determine analyte concentration using a smartphone-based colorimetric assay in a microfluidic device.
Materials:
- Smartphone with a dedicated colorimetry application (e.g., Appuente [6])
- Custom-designed cradle to hold the smartphone and microfluidic device [7] [4]
- Microfluidic Paper-based Analytical Device (μPAD) or polymer-based microfluidic chip
- Colorimetric reagents (e.g., enzymatic assays, redox indicators, pH indicators, or functionalized nanoparticles [4])
- Standard solutions of the target analyte
- Sample solutions
Method:
- Device Preparation: Functionalize the detection zone of the μPAD or microfluidic chip with the appropriate colorimetric reagent.
- Calibration: Introduce a series of standard solutions with known analyte concentrations into separate detection zones. Allow the colorimetric reaction to proceed to completion.
- Image Acquisition: Place the device in the custom cradle and use the smartphone application to capture images under consistent, controlled lighting conditions. The cradle ensures fixed alignment and distance [4].
- Color Space Conversion: The application automatically defines a Region of Interest (ROI) for each detection zone and converts the pixel values from RGB to a more perceptually uniform color space like HSV or CIE LAB* [4].
- Quantitative Analysis: The application fits the color intensity data (e.g., value in HSV or lightness in LAB*) from the calibration standards against concentration to generate a standard curve.
- Sample Measurement: Introduce the unknown sample into a separate detection zone, repeat steps 3 and 4, and use the standard curve to determine the analyte concentration.
Data Analysis: The limit of detection (LOD) and sensitivity are key performance metrics. Validation studies have shown high correlation (R² > 0.98) between smartphone image analysis and established software like ImageJ for parameters such as particle area and size [6].
Figure 1: Workflow for smartphone-based colorimetric detection.
Fluorescence Detection
Fundamental Principle
Fluorescence detection operates on the principle of photon absorption and re-emission at a longer wavelength. In this process, a fluorophore absorbs high-energy photons from an excitation light source, elevating electrons to an excited singlet state. Upon returning to the ground state, these electrons emit lower-energy photons (fluorescence) at a characteristic wavelength [5]. The difference between the peak excitation and emission wavelengths is known as the Stokes shift. The key parameters defining fluorescence include intensity, lifetime, polarization, and emission spectrum, each providing unique insights into the molecular environment and interactions.
Advanced Fluorescence Modalities
Several sophisticated fluorescence techniques enhance the capabilities of standard fluorescence intensity measurements:
- Fluorescence Polarization (FP): Utilizes polarized excitation light to measure the rotational diffusion of molecules. Binding of a small fluorescent ligand to a larger molecule slows its rotation, increasing the polarization of the emitted light, which is ideal for studying molecular interactions [5].
- Time-Resolved Fluorescence (TRF): Employs long-lived lanthanide fluorophores (e.g., Europium chelates) and measures emission after a delay from the excitation pulse. This minimizes short-lived background fluorescence, dramatically improving signal-to-noise ratios [5].
- Two-Photon Microscopy: A tissue-penetrating technique where a fluorophore is excited by the simultaneous absorption of two long-wavelength, low-energy photons. This reduces photodamage and allows for deeper imaging in biological tissues [8].
- Near-Infrared-II (NIR-II) Imaging: Uses fluorescent probes with emission in the 1000–1700 nm window, where tissue scattering and autofluorescence are minimal. This enables higher resolution and deeper penetration for in vivo imaging and drug tracking [8].
Experimental Protocol: Evanescent Wave Fluorescence Biosensing
Objective: To detect the binding of an analyte to a surface-immobilized ligand using evanescent wave-induced fluorescence.
Materials:
- Optical biosensor platform with integrated laser diode or LED source and waveguides [1] [3]
- Photomultiplier Tube (PMT), CCD, or CMOS sensor as detector [3]
- Sensor chip with functionalized surface (e.g., carboxymethylated dextran)
- Fluorescently-labeled analyte or secondary probe
- Running buffer and fluidics system for sample delivery
Method:
- Surface Functionalization: Immobilize the ligand (e.g., antibody, receptor) onto the sensor chip surface using appropriate chemistry (e.g., NHS/EDC for dextran chips [1]).
- Baseline Establishment: Flow running buffer over the sensor surface to establish a stable fluorescence baseline. The evanescent wave from the integrated waveguide only excites fluorophores within ~100-200 nm of the surface.
- Sample Injection & Binding: Inject the sample containing the fluorescently-labeled analyte. Binding events within the evanescent field result in a localized increase in fluorescence signal as the fluorophores are excited.
- Dissociation & Regeneration: Switch back to running buffer to monitor the dissociation of the complex. A regeneration solution may be used to remove bound analyte and prepare the surface for a new cycle.
- Data Analysis: The real-time binding curve (sensorgram) is analyzed to determine kinetic rate constants (kon, koff) and the equilibrium dissociation constant (KD) [1].
Data Analysis: For quantitative concentration analysis, the initial binding rate or steady-state response is measured and compared to a calibration curve. The evanescent nature of excitation effectively suppresses background fluorescence from the bulk solution, conferring high sensitivity.
Figure 2: Principle of evanescent wave fluorescence detection.
Interferometric Scattering and SPR-Based Detection
Fundamental Principle of Interferometry and SPR
Interferometric techniques, including Surface Plasmon Resonance (SPR) and reflectometric interference spectroscopy (RIfS), are powerful label-free methods that detect changes in the refractive index or optical thickness at a sensor surface [1]. SPR occurs when polarized light strikes a metal (typically gold) film at the interface of two media (e.g., glass and liquid) under specific conditions, generating charge density waves called surface plasmons [1]. This results in a drop in the intensity of the reflected light at a specific resonance angle. Any change in the mass on the metal surface, such as the binding of a biomolecule, alters the local refractive index and causes a measurable shift in the resonance angle [1]. Similarly, interferometric methods like RIfS monitor the interference pattern of light reflected from different layers of a sensor; binding events change the optical path length and thus the interference pattern.
Localized Surface Plasmon Resonance (LSPR)
A key variant, Localized SPR (LSPR), relies on metallic nanostructures (e.g., gold or silver nanoparticles) [1]. When incident light interacts with these nanostructures, it induces collective electron charge oscillations confined to the nanoparticle, leading to strong light absorption and scattering in the UV-visible range [1]. The LSPR phenomenon is highly sensitive to the local dielectric environment. A binding event on or near the nanoparticle surface causes a measurable shift in the LSPR absorption peak wavelength, enabling "wavelength-shift sensing" [1]. LSPR sensors are more adaptable for miniaturization and integration into portable devices compared to conventional SPR systems.
Experimental Protocol: LSPR-based Bioassay
Objective: To detect a specific analyte using the LSPR wavelength shift of functionalized gold nanoparticles.
Materials:
- Spectrometer or customized smartphone-based spectrophotometer [7]
- LSPR sensor chip (glass substrate with immobilized gold nanoparticles) or colloidal gold nanoparticles in solution [1]
- Ligand specific to the target analyte (e.g., antibody, DNA probe)
- Sample solutions containing the analyte
- Buffer solutions
Method:
- Sensor Functionalization: Immobilize the ligand onto the surface of the gold nanoparticles via thiol-gold chemistry or other suitable coupling methods.
- Baseline Spectrum: Acquire the extinction (absorbance + scattering) spectrum of the functionalized LSPR sensor to establish the initial peak resonance wavelength (λmax).
- Sample Exposure: Expose the sensor to the sample solution containing the analyte.
- Binding Measurement: Incubate to allow the analyte to bind to the surface-immobilized ligand. This binding event changes the local refractive index around the nanoparticles.
- Spectrum Acquisition: Acquire the final extinction spectrum and determine the new λmax.
- Quantification: The shift in the resonance wavelength (Δλ) is directly related to the analyte concentration and can be quantified against a calibration curve.
Data Analysis: The LSPR spectral shift is the primary readout. This method has been successfully applied for the detection of various targets, including viruses, toxins, and biomarkers, with demonstrated detection limits in the nanomolar to picomolar range [1]. Smartphone-based spectrometers have been shown to achieve resonant wavelength accuracy of up to 0.009 nm [7].
Figure 3: LSPR wavelength-shift sensing workflow.
Comparative Analysis of Detection Methods
Table 1: Performance comparison of optical detection methods in biosensing.
| Parameter | Colorimetric | Fluorescence | Interferometric/SPR |
|---|---|---|---|
| Principle | Absorption of light (Beer-Lambert) [4] | Emission of light after excitation [5] | Refractive index change [1] |
| Label Requirement | Often requires chromogenic label/dye | Requires fluorescent label [1] | Label-free [1] |
| Sensitivity (Typical LOD) | Moderate (µM–nM) | High (nM–pM) [8] | Very High (pM–fM) [1] |
| Multiplexing Potential | Moderate (spatial separation) | High (multiple colors/FRET) | High (SPR imaging) [1] |
| Hardware Complexity | Low (compatible with smartphones) [4] | Moderate to High (requires specific filters) | High (precision optics) |
| Cost | Low | Moderate to High | High |
| Primary Application Context | Point-of-care testing, rapid screening [6] [4] | Cellular imaging, drug tracking, high-sensitivity assays [8] | Kinetic binding studies, affinity characterization [1] |
The Scientist's Toolkit: Essential Research Reagents and Materials
Table 2: Key reagents and materials for optical detection experiments.
| Item | Function/Description | Example Applications |
|---|---|---|
| Gold Nanoparticles (AuNPs) | Plasmonic nanoparticles for LSPR sensing and colorimetric labels due to high extinction coefficients [1] [4]. | LSPR bioassays, colorimetric aggregation assays. |
| Nile Red (NR) Dye | Hydrophobic fluorescent dye that adsorbs to plastics, used for staining and detecting microplastics [6]. | Fluorescent identification and counting of microplastics. |
| Carboxymethylated Dextran Matrix | Hydrogel matrix for immobilizing ligands on sensor surfaces via NHS/EDC chemistry [1]. | SPR and BLI sensor chips for biomolecular interaction analysis. |
| Quantum Dots (QDs) | Semiconductor nanocrystals with size-tunable fluorescence and high brightness; used as fluorescent labels [8] [3]. | Highly multiplexed assays, long-term cell tracking. |
| NIR-II Fluorophores | Fluorescent probes emitting in the 1000–1700 nm range for deep-tissue imaging with reduced scattering [8]. | In vivo drug tracking, deep-tissue diagnostics. |
| Lanthanide Chelates (e.g., Eu³⁺) | Long-lifetime fluorophores for Time-Resolved Fluorescence (TRF), minimizing background autofluorescence [5]. | TRF-based immunoassays (e.g., DELFIA), high-throughput screening. |
| Microfluidic Chips (μPAD/PDMS) | Miniaturized platforms for automating sample handling and reaction containment [4] [3]. | Lab-on-Chip diagnostics, point-of-care testing devices. |
| Photomultiplier Tube (PMT) | Highly sensitive light detector that multiplies incident photons via electron cascade, used in many plate readers [3] [5]. | Detecting low-intensity fluorescence and luminescence signals. |
The convergence of smartphone technology with biosensing has created a paradigm shift in point-of-care (POC) diagnostics, environmental monitoring, and food safety analysis. This integration effectively transforms ubiquitous mobile devices into portable, sophisticated analytical platforms, making laboratory-grade sensing accessible outside traditional settings [9]. The core smartphone components—high-resolution cameras, powerful processors, and versatile connectivity options—serve as the foundation for these biosensing systems, enabling the detection of a wide range of analytes from pathogens and proteins to metabolites and toxins [10] [11].
Framed within the broader principles of optical detection methods in smartphone-based Lab-on-Chip (LoC) research, this technical guide explores how smartphones interact with optical biosensors. These systems leverage fundamental phenomena including colorimetry, fluorescence, chemiluminescence, and label-free detection methods such as surface plasmon resonance (SPR) and photonic crystal (PC) sensing [10]. The proliferation of smartphones, with an estimated 51% of the ~7.5 billion mobile phones in use classified as "smart" as of 2017, provides an unprecedented infrastructure for deploying diagnostic technology [10]. This whitepaper details the core technical principles, methodologies, and material requirements for developing and implementing smartphone-based biosensing platforms.
Core Smartphone Components in Biosensing
The functionality of smartphones as biosensors hinges on three primary subsystems: the camera as a detector, the processor for data analysis, and connectivity for data transmission.
Camera as a Optical Detector
The smartphone camera, typically a complementary metal-oxide-semiconductor (CMOS) sensor, functions as a versatile spectrometer and imager. It captures optical signals—changes in color, intensity, or wavelength—generated by biochemical reactions on sensor surfaces or within assay platforms [9] [10] [11]. For instance, in colorimetric assays, the camera captures images of color changes, which are then converted into quantitative values in color spaces like RGB (Red, Green, Blue) or HSV (Hue, Saturation, Value) [12]. Advanced implementations use a cradle containing optical components like diffraction gratings to allow the onboard camera to function as a high-resolution spectrometer, capable of measuring shifts in wavelength resulting from biological adsorption onto a sensor surface [10] [13]. This system can perform as accurately as a large laboratory spectrophotometer at a fraction of the cost [13].
Processor as a Data Analyzer
The smartphone's central processing unit (CPU) provides the computational power for real-time data processing and analysis. This includes running algorithms for image analysis, color space conversion, spectral data interpretation, and concentration interpolation from calibration curves [9] [14]. The processor executes the software that drives the assay, controls hardware components (e.g., excitation sources), and delivers a user-friendly interface, making sophisticated diagnostic tools accessible to non-specialists [15]. The integration of artificial intelligence (AI) and machine learning (ML) algorithms further enhances the capability for complex pattern recognition and multi-analyte analysis [9] [14].
Connectivity for Data Transmission
Smartphones offer multiple integrated options for data transmission, which is crucial for telemedicine and networked health care systems.
- Wireless Peripherals (Wi-Fi and Bluetooth): These are widely used for communication with external sensors and for transmitting data to cloud-based storage or monitoring systems [14]. Bluetooth is valued for its high compatibility across phone models, while Wi-Fi offers greater bandwidth and integration with existing internet infrastructure [14].
- Near-Field Communication (NFC): This technology is particularly promising for contactless biosensing due to its capability for low-power data transfer and even wireless powering of simple circuits [14] [16].
- Wired Peripherals (USB and Audio Jack): The USB port and audio jack provide stable, practical connections for powering external sensor modules and transmitting data, albeit with the inconvenience of physical cables [14].
Optical Detection Modalities and Experimental Protocols
The following table summarizes the primary optical detection modalities used in smartphone-based biosensing.
Table 1: Key Optical Detection Modalities in Smartphone-Based Biosensing
| Detection Modality | Principle | Typical Assay Format | Key Advantages | Inherent Challenges |
|---|---|---|---|---|
| Colorimetric | Measures change in light absorption/reflectance due to color change [12]. | µPADs, lateral flow assays (LFA), microfluidic chips [15] [14]. | Simplicity, rapid response, naked-eye qualitative readout [12]. | Poor accuracy in variable light, requires clear samples [12]. |
| Fluorescent | Measures light emission from an excited substance [12]. | Microfluidic chips, molecular beacon FRET assays [10] [12]. | High sensitivity and specificity [12]. | Background interference; requires excitation sources/filters [10] [12]. |
| Chemiluminescent | Measures light radiation from chemical reactions [12]. | ELISA, immunodetection assays. | High signal-to-noise ratio, no excitation light needed [12]. | Low luminescence intensity, can be time-consuming [12]. |
| Label-Free (e.g., SPR/PC) | Measures shift in optimal optical coupling due to analyte adsorption [10]. | Photonic crystal (PC) biosensors. | Label-free, real-time monitoring, high sensitivity [10]. | Requires precise optical alignment (e.g., cradle) [10] [13]. |
Detailed Protocol: On-Chip Colorimetric Detection of Pathogen RNA
The following workflow describes a protocol for the label-free detection of Cryptosporidium RNA using a smartphone-integrated, on-chip colorimetric platform [17].
Principle: Thiolated oligonucleotide probes specific to target RNA sequences are immobilized on gold nanoparticles (AuNPs). In the presence of the complementary RNA, hybridization occurs, leading to AuNP aggregation. This aggregation causes a localized surface plasmon resonance (LSPR) shift, resulting in a visible color change from red to blue, which is quantified by a smartphone camera [17].
Figure 1: Workflow for smartphone-based colorimetric RNA detection.
Materials and Reagents:
- Gold Nanoparticles (AuNPs): 20 nm diameter, citrate-capped [17].
- Thiolated Oligonucleotides: Designed to be complementary to adjacent sequences on the target Cryptosporidium RNA [17].
- Buffer Solutions: Sodium phosphate buffer (for probe immobilization), sodium chloride (SDS) solution.
- DL-Dithiothreitol (DTT): Used to reduce disulfide bonds in thiolated oligonucleotides before conjugation [17].
- Illustra NAP-5 Columns: For purifying probe-conjugated AuNPs from excess reagents [17].
- Microfabricated Chip: The substrate for performing the assay.
- 3D-Printed Holder: A portable assembly that holds the smartphone and chip in fixed alignment for consistent imaging [17].
Experimental Procedure:
- Probe Preparation: Reduce the disulfide bonds of the thiolated oligonucleotides in DTT solution for 1 hour. Purify the reduced oligonucleotides using a NAP-5 column [17].
- AuNP Functionalization: Incubate the purified thiolated oligonucleotides with the 20 nm AuNPs for 16-24 hours to allow the thiol groups to bind to the gold surface. Pass the solution through a NAP-5 column to remove unbound oligonucleotides [17].
- Assay Execution: Mix the functionalized AuNPs with the extracted RNA sample. Incubate the mixture for 5-30 minutes at room temperature to allow for hybridization and aggregation [17].
- Image Acquisition: Pipette the mixture into the wells of the microfabricated chip placed in the 3D-printed holder. Use the smartphone camera to capture an image of the chip under consistent lighting conditions, ideally within a dark box to minimize ambient light interference [17].
- Data Analysis: A custom application on the smartphone processes the captured image. It quantifies the color change by analyzing the RGB or HSV values in a defined region of interest. The concentration of the target RNA is determined by interpolating the color value against a pre-established calibration curve from standards of known concentration [17].
This platform demonstrated a wide linear response (5–100 µM) and a detection limit of 5 µM for Cryptosporidium RNA, showing high specificity against non-complementary RNA sequences [17].
Detailed Protocol: Wireless Electrochemiluminescent (ECL) Biosensor
This protocol describes a fully wireless biosensor where a smartphone powers the sensing chip and detects the emitted light, positioning it as a potent Internet of Things (IoT) tool [16].
Principle: An electrode chip, without integrated circuits, receives power via electromagnetic induction from the smartphone. This power induces an electrochemiluminescence reaction (e.g., of luminol) on the printed electrode. The resulting luminescence is quantitatively detected by the smartphone's high-sensitivity CMOS camera [16].
Materials and Reagents:
- Printed Electrode Chip: Mass-producible, low-cost electrode.
- Enzyme Solution: Glucose oxidase (GOD) encapsulated in a chitosan polymer matrix to maintain high activity [16].
- ECL Substrate: Luminol solution.
- Circuit Components: Inductance and capacitance components that resonate with the transmission frequency, and a diode for rectification [16].
Experimental Procedure:
- Enzyme Immobilization: Immobilize the glucose oxidase enzyme in a chitosan polymer matrix on the counter electrode of the printed chip. (Immobilization on the working electrode was found to suppress luminescence) [16].
- Wireless Power Activation: Bring the electrode chip, containing a drop of the sample mixed with ECL substrate, near the smartphone. The smartphone wirelessly powers the electrode via electromagnetic induction through a resonant circuit on the chip, inducing the ECL reaction [16].
- Signal Detection and Analysis: The light generated from the ECL reaction on the electrode is captured by the smartphone's camera. Open-source image analysis software is used to quantitatively evaluate the images, determining the concentration of the target analyte (e.g., glucose) [16].
This system was successfully tested with human serum and artificial sweat samples, demonstrating its potential for real-world POC applications [16].
The Scientist's Toolkit: Essential Research Reagents and Materials
The development and execution of smartphone-based biosensing experiments require a suite of specialized materials and reagents. The following table catalogs key components.
Table 2: Essential Research Reagents and Materials for Smartphone-Based Biosensing
| Item Category | Specific Examples | Function in the Biosensing Platform |
|---|---|---|
| Nanomaterials | Gold Nanoparticles (AuNPs) [17], Quantum Dots [12] | Act as signal generators or reporters; AuNPs exhibit LSPR shifts for colorimetric detection [17]. |
| Biorecognition Elements | Thiolated Oligonucleotides [17], Enzymes (e.g., Glucose Oxidase) [16], Antibodies [9], Bacteriophages [9] | Provide high specificity and selectivity by binding to the target analyte (DNA, RNA, proteins, bacteria) [9] [17]. |
| Substrates & Chips | Microfluidic Paper-Based Analytical Devices (µPADs) [15], Microfluidic Chips [14], Photonic Crystals (PC) [10], Printed Electrodes [16] | Serve as the platform for housing the assay, facilitating fluid control, and serving as the transducer surface. |
| Polymers & Chemicals | Chitosan [16], Dextran Sulfate [17], DL-Dithiothreitol (DTT) [17], Luminol [16] | Used for enzyme immobilization, assay buffers, signal generation, and probe preparation. |
| Optical Components | Diffraction Gratings [10], Emission/Excitation Filters [10] [12], LEDs [12], 3D-Printed Cradles & Dark Boxes [13] [17] | Constitute the external hardware that interfaces with the smartphone to create a controlled optical environment for precise measurements. |
Current Challenges and Future Perspectives
Despite significant advancements, the transition of smartphone-based biosensors from research laboratories to widespread commercial adoption faces several hurdles.
A critical analysis of patent applications reveals a sharp decline after 2016, suggesting challenges in technology transfer and implementation with real samples [15]. Technical limitations include the reproducibility and repeatability of assays, particularly those using paper-based substrates, and the complexity of miniaturizing optical systems while maintaining robustness [15]. Furthermore, obtaining regulatory approvals for clinical use and achieving seamless end-user adoption outside research settings remain significant barriers [15].
Future development will likely focus on several key areas. Multiplexed detection, or the simultaneous measurement of multiple biomarkers in a single test, is crucial for accurate diagnosis of complex diseases like cancer and cardiovascular conditions [14]. The deep integration of AI and cloud computing will enhance data analysis, enable personalized health monitoring, and support networked healthcare systems [9] [14]. Finally, the creation of self-contained, fully wireless systems, such as the wireless ECL biosensor, will be pivotal in advancing IoT biosensors for effortless POC testing [16]. As these technologies mature, smartphone-based biosensing platforms are poised to become indispensable tools in transforming global healthcare, environmental safety, and food security landscapes.
The convergence of microfluidics and smartphone technology is forging a new paradigm in portable molecular analysis. These integrated systems are poised to transform point-of-care testing (POCT), environmental monitoring, and personalized medicine by making sophisticated laboratory capabilities accessible in resource-limited settings [18] [19]. The core innovation lies in harmonizing microfluidic precision with the smartphone's ubiquitous presence, computational power, and advanced sensors [20]. This technical guide examines the fundamental components that constitute these platforms, framed within the broader context of optical detection methods in smartphone-based lab-on-chip (LoC) research. For researchers and drug development professionals, understanding these synergistic elements is crucial for developing robust, field-deployable diagnostic tools that transcend traditional laboratory boundaries.
Core Microfluidic Platforms for Sample Handling
Microfluidic chips form the analytical heart of smartphone-based LoC systems, responsible for precise fluid manipulation and housing biological or chemical reactions. The design and material selection for these chips are paramount, dictating the platform's functionality, cost, and suitability for specific applications.
Table 1: Comparison of Microfluidic Chip Substrates and Their Properties
| Material | Key Advantages | Limitations | Common Fabrication Methods | Ideal Use Cases |
|---|---|---|---|---|
| Polydimethylsiloxane (PDMS) | Excellent transparency, flexibility, gas permeability [20] | Susceptible to adsorption of biomolecules [20] | Soft lithography, molding [20] | Prototyping, biological applications [20] |
| Polymethylmethacrylate (PMMA) | High durability, low cost, chemical resistance [20] | Lower optical clarity than glass, limited thermal stability [20] | Injection molding, laser cutting [20] | Disposable environmental & agricultural sensors [20] |
| Paper | Extremely low cost, capillary-driven flow (pump-free) [20] | Sensitive to environmental humidity, less durable [20] | Wax printing [21] | Rapid diagnostics, lateral flow assays [20] |
| Glass | Superior optical clarity, chemical stability [20] | High cost, difficult fabrication [20] | Etching, bonding [20] | High-precision fluorescence assays [20] |
| 3D-Printable Resins | Rapid prototyping, complex 3D geometries [22] | May require surface treatment for hydrophilicity [22] | Stereolithography (SLA) [22] | Custom, monolithic auto-mixing devices [22] |
Three dominant fluidic paradigms have emerged in smartphone-based systems:
- Lateral and Vertical Flow Assays: These pump-free platforms leverage capillary action to transport samples. Lateral flow devices (e.g., similar to pregnancy tests) are cost-effective but can have limitations in sensitivity and multiplexing capability [18]. A key improvement is the Vertical Flow Assay (VFA), which utilizes a porous membrane with separated spots for simultaneous multiplexed analysis, with results readable by a smartphone [18].
- Microchannel Capillary Flow Assays (MCFA): These chips use microchannels with precisely engineered geometry and surface properties to control fluid flow without external pumping [18]. This design allows for more complex fluidic pathways than paper-based devices while maintaining a passive flow system.
- Active Microfluidic Chips: For procedures requiring precise fluidic control, such as nucleic acid amplification, chips can integrate miniature active components. Examples include finger-operated pumps [18], passive vacuum pumps [18], and even integrated heaters for temperature control [18]. Gou et al. developed a platform that integrated thermal cycling control with on-chip digital polymerase chain reaction for DNA quantification [18].
Optical Detection Modalities and Smartphone Integration
The smartphone camera serves as the primary detector, leveraging its advanced complementary metal-oxide-semiconductor (CMOS) sensor to capture optical signals generated within the microfluidic chip. The choice of optical method depends on the target analyte and required sensitivity.
Primary Optical Detection Methods
- Colorimetry: This method detects the color change of a reaction, often quantified by the smartphone camera in the RGB (Red, Green, Blue) color space. It is widely used due to its simplicity. For instance, a 3D-printed auto-mixing chip was used for a colorimetric hemoglobin assay, where RGB values were converted to a device-independent color space (CIE Lab*) for concentration analysis [22]. Similarly, a paper microfluidic device with a colorimetric reagent (RBCl) enabled smartphone-based detection of copper ions (Cu²⁺) [21].
- Fluorescence: Fluorescence-based detection offers higher sensitivity and specificity compared to colorimetry. It requires an external light source (e.g., an LED) to excite the fluorescent labels and an emission filter to block the excitation light, allowing only the emitted signal to reach the camera [18] [19].
- Chemiluminescence: This method detects light emitted as a result of a chemical reaction, eliminating the need for an excitation light source. This simplifies the optical setup, as it only requires the camera to be aligned with the reaction chamber to capture the emitted light [23].
- Brightfield Microscopy: This is used for imaging micro-scale objects like cells or parasites. It can be implemented in a lens-free configuration, using the sample's shadow on the sensor, or with added lenses for magnification [18].
Supporting Hardware Attachments
To interface the microfluidic chip with the smartphone camera and enable these detection modalities, custom hardware attachments are essential. These components are increasingly fabricated using 3D printing, which allows for rapid prototyping and customization to specific smartphone and chip geometries [22].
- Lenses: External lenses are often added to the smartphone's native camera to provide the magnification needed to resolve microscopic features or small reaction zones. These can range from simple clip-on magnifiers to more complex, aligned lens systems [18] [22].
- Light Sources: Controlled illumination is critical. Light-emitting diodes (LEDs) are the standard source due to their small size, low power consumption, and spectral variety. They provide uniform brightfield illumination [18] or specific wavelengths for fluorescence excitation [19].
- Housings: A 3D-printed housing or adapter is the structural backbone that ensures precise and reproducible alignment of the microfluidic chip, optical components, and the smartphone camera. This is vital for obtaining quantitative and reliable data [22] [21].
Table 2: Quantitative Performance of Smartphone-Based Detection Systems
| Detection Target | Optical Method | Microfluidic Platform | Reported Limit of Detection (LOD) | Assay Time | Citation |
|---|---|---|---|---|---|
| Copper Ions (Cu²⁺) | Colorimetry | Paper-based device | 1.51 ng/mL | < 2 minutes | [21] |
| Blood Hemoglobin | Colorimetry | 3D-printed auto-mixer | Clinical concordance (a.u.c. = 0.97) | ~1 second mixing | [22] |
| HIV | Colorimetry (Lateral Flow) | Lateral Flow Strip | 97.8% Sensitivity, 100% Specificity | Rapid test | [18] |
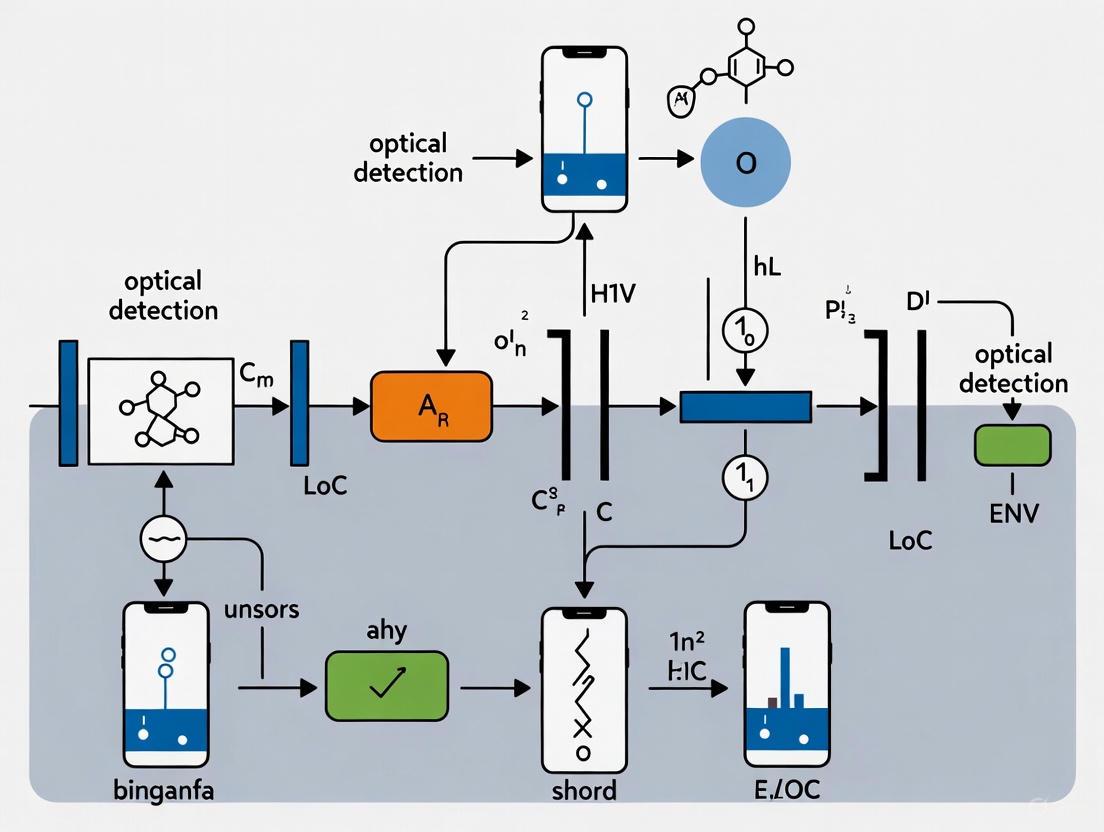
The following workflow diagram illustrates the integration of these components and the process of conducting an analysis.
Figure 1: Integrated Workflow of a Smartphone-Based LoC Platform. The process involves both the physical integration of hardware components and the sequential steps of the analytical assay.
Experimental Protocol: Colorimetric Hemoglobin Detection
The following detailed protocol, adapted from a published study, exemplifies the application of the components discussed above to create a functional quantitative test [22].
Objective: To perform a rapid, quantitative colorimetric assay for blood hemoglobin (Hgb) levels using a 3D-printed auto-mixing microfluidic chip and a smartphone reader.
Materials and Reagents:
- 3D-Printed Microfluidic Auto-mixer: Fabricated using a consumer-grade 3D printer (e.g., D3 ProJet 1200) and VisiJetFTX Clear resin. The chip design features microstructures optimized via computational fluid dynamic (CFD) simulation for rapid capillary-driven mixing [22].
- Smartphone & Attachment: An Android smartphone running a custom color-scale analytical application, housed in a 3D-printed enclosure that aligns the phone's camera with a 5x magnification gel lens and the microfluidic chip's view window [22].
- Chemical Reagents: 3,3',5,5'-Tetramethylbenzidine (TMB) and hydrogen peroxide (H₂O₂) for the hemoglobin-catalyzed oxidation-reduction reaction [22].
- Blood Sample: 5 μL of finger-prick blood, diluted 10x [22].
- Ethylene glycol/KOH solution: For post-printing surface treatment to render the 3D-printed chip hydrophilic [22].
Procedure:
- Chip Fabrication and Preparation:
- Design the microfluidic auto-mixer using CAD software (e.g., AutoCAD 360) and 3D print it as a monolithic device using clear resin.
- Clean the printed device with isopropyl alcohol and flush the channels with compressed air.
- Treat the chip by soaking it in a 1.82 M KOH solution in ethylene glycol at 55°C for 2 hours. This ethylene glycol chemistry creates a hydrophilic surface to facilitate capillary action [22].
- Assay Execution:
- Introduce the 5 μL diluted blood sample into the chip's input well.
- The capillary force and designed microstructures will automatically draw the sample into the mixing zone, combining it with pre-loaded or subsequently added TMB/H₂O₂ reagents. Mixing is achieved in approximately 1 second [22].
- Hemoglobin catalyzes the reaction, producing a color change in the view window proportional to its concentration.
- Smartphone Readout and Analysis:
- Place the chip into the dedicated slot in the 3D-printed smartphone housing, ensuring the view window is aligned with the camera and lens.
- Launch the custom color-scale analytical app.
- Capture an image of the view window. The app will define a region of interest (ROI) and extract the average RGB pixel values.
- The app internally converts the RGB values to CIE Lab* color space values, which are correlated to hemoglobin concentration via a pre-established calibration curve stored in the app [22].
Validation: In a clinical training set (n=22), this iPOC3D system demonstrated consistent measurements with a high diagnostic accuracy (area under curve, a.u.c. = 0.97) compared to a standard clinical hematology analyzer [22].
Essential Research Reagent Solutions
The development and operation of these integrated systems rely on a suite of specialized reagents and materials. The following table details key components used in the featured experiments and the broader field.
Table 3: Key Research Reagents and Materials for Smartphone LoC Systems
| Reagent / Material | Function / Role | Example Application |
|---|---|---|
| 3,3',5,5'-Tetramethylbenzidine (TMB) | Chromogenic substrate; produces color change upon oxidation catalyzed by an enzyme or catalyst like hemoglobin. | Colorimetric detection of blood hemoglobin [22]. |
| Rhodamine B Derivative (RBCl) | Colorimetric and fluorescent chemodosimeter; specific structural transition (ring-opening) upon binding to target ions. | Selective detection of Copper ions (Cu²⁺) on paper microfluidics [21]. |
| VisiJetFTX Clear Resin | Photopolymer resin for high-resolution 3D printing; enables fabrication of transparent, monolithic microfluidic devices. | Production of 3D-printed auto-mixing chips [22]. |
| Polydimethylsiloxane (PDMS) | Elastomeric polymer for soft lithography; gas permeable and optically transparent. | Fabrication of flexible microfluidic chips for cell culture and analysis [20]. |
| Ethylene Glycol / KOH Solution | Surface treating agent; confers hydrophilic properties to otherwise hydrophobic 3D-printed surfaces. | Post-printing treatment of resin-based chips to enable capillary flow [22]. |
| Whatman Chromatography Paper | Porous cellulose matrix; serves as a pump-free platform for fluid transport via capillary action. | Substrate for paper microfluidic devices [21]. |
The integration of microfluidics, custom optics (LEDs, lenses), and 3D-printed attachments with smartphones creates a powerful and versatile platform for optical detection outside the conventional laboratory. The synergy between these components—where the microfluidic chip handles the assay chemistry, the optical hardware enables signal transduction, and the smartphone provides computation, control, and connectivity—is the cornerstone of this technology. As evidenced by the quantitative performance in detecting analytes from hemoglobin to metal ions, these systems are maturing into reliable tools for researchers and clinicians. Future advancements will likely be driven by improvements in AI-powered image analysis [18], the development of even more robust and inexpensive materials [20], and a focus on multiplexing capabilities to enable comprehensive diagnostic panels at the point of need. For the field to fully translate from research prototypes to widespread practical application, ongoing efforts must focus on standardizing validation protocols and ensuring user-friendly design for non-experts.
Point-of-need (PON) analysis represents a paradigm shift in chemical and biological testing, moving traditional laboratory processes directly to the location where information is needed. This transition is fundamentally enabled by the convergence of miniaturized technologies and smartphone-based detection platforms that align with the core principles of Green Analytical Chemistry (GAC). Within the broader context of a thesis on optical detection methods in smartphone-based lab-on-a-chip (LoC) research, this whitepaper examines how portability, cost-effectiveness, and environmental sustainability are interconnected advantages that reinforce one another in modern analytical system design.
The drive toward PON analysis stems from several critical needs in healthcare, environmental monitoring, and food safety. Residents of rural and remote communities, representing an underserved 45% of the global population, often lack access to centralized laboratory facilities [19]. Furthermore, applications such as disease outbreak investigation, environmental contamination assessment, and personalized medicine demand analytical capabilities that are rapid, decentralized, and accessible without substantial financial barriers [19]. Smartphone-based LoC systems address these needs by leveraging the ubiquitous presence of mobile technology while simultaneously reducing the environmental impact of traditional analytical methods.
Technological Foundations of Smartphone-Based Point-of-Need Analysis
Smartphone as an Integrated Analytical Platform
Modern smartphones provide a uniquely integrated package of technologies that enable sophisticated chemical and biological analysis without the need for extensive custom engineering. These devices incorporate multiple sensing capabilities, processing power, and connectivity features that make them ideal foundations for PON diagnostic systems [19].
The camera system serves as the primary optical detection component, with specifications that have advanced dramatically in recent years. As illustrated in Table 1, smartphone camera capabilities now rival those of specialized scientific instrumentation in many applications, providing sufficient resolution and sensitivity for various colorimetric, fluorometric, and luminescence detection methods [19].
Table 1: Key Smartphone Features Enabling Point-of-Need Analysis
| Smartphone Feature | Technical Specifications | Analytical Function |
|---|---|---|
| Camera System | 12-108 MP sensors; ƒ/1.5-2.4 aperture; 4K video recording | Optical detection (absorbance, fluorescence, microscopy) |
| Processing Power | Multi-core CPUs (>2.8 GHz); 4-8 GB RAM | Real-time data processing and analysis |
| Connectivity | 5G, Wi-Fi 6, Bluetooth 5.2 | Data transmission and cloud integration |
| Sensors | Accelerometer, gyroscope, magnetometer, GPS | Sample orientation, flow timing, location tagging |
| Battery | 3000-5000 mAh capacity | Portable power for field analysis |
| Display | 6-7 inch OLED/IPS LCD (>450 ppi) | Result visualization and user interface |
The global penetration of smartphone technology creates an unprecedented opportunity for deploying analytical capabilities at scale. With approximately 54% of the world's population owning smartphones and mobile networks available to 95% of people, the infrastructure for deploying PON analysis already exists [19]. This existing distribution network significantly reduces the barriers to implementing analytical systems in resource-limited settings.
Principles of Optical Detection in Smartphone-Based LoC
Optical detection methods form the cornerstone of most smartphone-based analytical systems due to the sophisticated camera technology available in these devices. The fundamental principle involves coupling microfluidic or paper-based analytical devices with the smartphone's camera to capture optical signals that correlate with analyte concentration. Common approaches include:
- Colorimetric Detection: Measuring color intensity changes from chemical reactions using ambient light or integrated LED flashes
- Fluorescence Detection: Quantifying emitted light from labels or native fluorescence using additional excitation sources
- Chemiluminescence Detection: Capturing light emitted from chemical reactions without requiring an excitation source
- Bright-field Microscopy: Imaging samples with additional lens attachments for cellular or particle analysis
The experimental workflow for developing smartphone-based optical detection systems typically follows a structured approach, as detailed below:
Diagram 1: Smartphone assay development workflow.
For colorimetric detection, a typical protocol involves:
- Device Fabrication: Creating microfluidic channels (~50-200 µm wide) in PDMS using soft lithography or using paper-based microfluidic substrates
- Sample Introduction: Applying liquid sample (1-50 µL) to the device inlet via capillary action or pipetting
- Reaction Incubation: Allowing sufficient time (30-300 seconds) for color development at controlled temperature
- Image Capture: Positioning the smartphone camera at a fixed distance (5-15 cm) with consistent lighting conditions
- Image Analysis: Converting RGB values to grayscale or hue-saturation-intensity models to quantify color intensity
For fluorescence-based assays, the methodology requires:
- Excitation Source: Integrating LEDs (365-470 nm) with appropriate bandpass filters
- Emission Filtering: Placing emission filters between the sample and camera to block scattered excitation light
- Signal Capture: Using the smartphone camera in manual mode with fixed ISO, exposure time, and focus settings
- Background Subtraction: Applying image processing algorithms to remove background fluorescence
- Intensity Quantification: Correlating pixel intensity with analyte concentration using calibration standards
Advantages of Point-of-Need Analysis Systems
Portability and Accessibility
The miniaturization of analytical systems represents one of the most significant advantages for PON testing. Traditional laboratory instrumentation often requires dedicated space, stable benchtops, and controlled environments, whereas smartphone-based LoC devices can be transported and deployed in virtually any setting. This portability is achieved through several key technological developments:
- Microfluidics and MEMS Technology: Microelectromechanical systems (MEMS) combine mechanical parts, sensors, actuators, and electronics on a common silicon substrate, creating complex machines with sizes in the micrometer range [24]. These systems enable complete analytical processes to be performed in devices that fit in the palm of the hand.
- Miniaturized Detection Systems: The integration of optical components with microfluidic devices eliminates the need for bulky microscopes or spectrophotometers. Smartphone cameras, when properly configured with simple optical attachments, can achieve detection limits comparable to benchtop systems for many applications.
- Passive Fluid Control: The development of capillary-driven microfluidic systems removes the requirement for external pumps or power sources, further enhancing portability [19].
The relationship between portability and analytical performance in smartphone-based systems involves careful balancing of multiple engineering parameters, as shown in Diagram 2.
Diagram 2: Portability and performance engineering balance.
The accessibility benefits of portable PON systems extend beyond mere convenience. In healthcare applications, these devices enable rapid screening and diagnosis in primary care settings, remote communities, and home-based testing environments. Studies have demonstrated the effectiveness of smartphone-based detection for paediatric ocular diseases, with 33 included studies involving 16,015 participants showing comparable accuracy to conventional methods for conditions including refractive errors, strabismus, and retinopathy of prematurity [25]. Similar approaches have been applied to infectious disease testing, environmental monitoring, and food safety assessment.
Cost-Effectiveness and Economic Viability
The economic advantages of smartphone-based PON analysis systems operate at multiple levels, from initial capital investment to operational expenses. The foundation of this cost-effectiveness stems from leveraging the existing consumer electronics market, which provides sophisticated technology at a fraction of the cost of specialized scientific equipment.
- Hardware Cost Reduction: The annual smartphone market exceeds 1.3 billion units valued at $500 billion USD, creating economies of scale that dramatically reduce manufacturing costs [19]. A smartphone with capable camera and processing features costs between $100-1200 USD, compared to specialized analytical instruments that often range from $10,000 to $100,000.
- Elimination of Peripheral Equipment: Traditional microfluidic systems often require peripheral equipment for operation, including "pumps, pressure generators, power supplies, voltage sequencers, temperature controllers, light sources, microscopes, photodetectors, potentiostats, and other hardware" [19]. Smartphone-based systems integrate or eliminate many of these components.
- Reduced Reagent Consumption: Microfluidic LoC devices typically require smaller sample and reagent volumes (microliter range compared to milliliters in conventional systems), leading to significant cost savings, particularly for expensive biological reagents.
Table 2: Cost Comparison of Analytical Approaches
| Cost Factor | Traditional Laboratory Analysis | Smartphone PON System |
|---|---|---|
| Initial Instrument Cost | $10,000 - $100,000 | $100 - $1,200 (smartphone) |
| Per Test Consumable Cost | $5 - $100 | $0.50 - $10 |
| Sample Volume Requirements | 0.5 - 10 mL | 1 - 100 µL |
| Personnel Requirements | Trained technical staff | Minimal training required |
| Maintenance Costs | High (service contracts, calibration) | Low (consumer electronics warranty) |
| Result Turnaround Time | Hours to days | Minutes to hours |
From an implementation science perspective, cost-effectiveness analysis (CEA) provides a framework for evaluating the trade-offs decision makers face when considering alternative courses of action for implementing public health strategies [26]. The RE-AIM framework (Reach, Effectiveness, Adoption, Implementation, Maintenance) offers a structured approach to evaluating these economic factors, particularly regarding scalability and sustainability [26].
The economic value of PON systems extends beyond direct cost savings to include opportunity cost reductions associated with faster decision-making. In clinical settings, rapid diagnosis enables timely treatment interventions that can improve outcomes and reduce overall healthcare costs. In environmental monitoring, immediate detection of contaminants allows for quicker remediation responses, potentially preventing more widespread contamination.
Green Analytical Chemistry Integration
The alignment between PON analysis and Green Analytical Chemistry principles represents a synergistic relationship that enhances both environmental sustainability and analytical efficiency. GAC is defined as "the optimization of analytical processes to ensure they are safe, nontoxic, environmentally friendly, and efficient in their use of materials, energy, and waste generation" [27]. Smartphone-based PON systems advance these goals through several mechanisms:
- Miniaturization and Waste Reduction: The small dimensions of LoC devices directly reduce consumption of reagents and samples, with volumes typically in the microliter range compared to milliliters in conventional methods [28]. This miniaturization correspondingly decreases waste generation, addressing the first principle of green chemistry: waste prevention [29].
- Solvent Replacement and Elimination: Traditional analytical methods often rely on large quantities of organic solvents for extraction and separation. GAC approaches replace these with bio-based solvents, ionic liquids, and deep eutectic solvents that are less toxic and more biodegradable [28] [29]. In some cases, solvent-free techniques entirely eliminate this waste stream.
- Energy Efficiency: Smartphone-based detection typically requires less power than benchtop instrumentation. While laboratory instruments may consume hundreds of watts, a smartphone operates at 5-10 watts, with additional efficiency gains from the elimination of peripheral equipment [29].
The relationship between GAC principles and smartphone-enabled PON technologies creates a self-reinforcing cycle of improvement, as illustrated in Diagram 3.
Diagram 3: GAC and PON technology synergy.
Several assessment tools have been developed to quantify the greenness of analytical methods, including:
- NEMI (National Environmental Methods Index): Provides a simple pictogram indicating whether a method meets basic green criteria [27]
- GAPI (Green Analytical Procedure Index): Offers a comprehensive color-coded evaluation of the entire method lifecycle [27]
- AGREE (Analytical GREEnness): Provides a holistic assessment based on all 12 GAC principles [27]
When applied to smartphone-based PON systems, these tools typically demonstrate superior environmental performance compared to traditional laboratory methods, particularly in categories related to reagent consumption, waste generation, and energy requirements.
Research Reagent Solutions and Materials
The development and implementation of smartphone-based PON analysis requires specialized materials and reagents that enable miniaturized, sensitive detection while maintaining alignment with green chemistry principles. Table 3 outlines key research reagent solutions and their functions in these analytical systems.
Table 3: Essential Research Reagent Solutions for Smartphone-Based PON Analysis
| Material/Reagent | Function | Green Alternatives |
|---|---|---|
| Polydimethylsiloxane (PDMS) | Microfluidic device fabrication; optical clarity; gas permeability | Biodegradable polymers; paper substrates |
| Nitrocellulose Membrane | Lateral flow assays; protein immobilization | Modified cellulose papers |
| Gold Nanoparticles | Colorimetric labels; surface plasmon resonance | Carbon nanoparticles; fluorescent nanocrystals |
| Deep Eutectic Solvents (DES) | Green extraction media; non-toxic and biodegradable | Bio-based solvents; supercritical CO₂ |
| Ionic Liquids (ILs) | Green solvents; stationary phases in separations | Switchable solvents; natural deep eutectic solvents |
| Enzyme Substrates | Signal generation in bioassays (e.g., chromogenic/fluorogenic) | Natural product-derived substrates |
| Molecularly Imprinted Polymers (MIPs) | Synthetic recognition elements; sample preparation | Biopolymer-based recognition elements |
| Quantum Dots | Fluorescent labels; broad excitation, narrow emission | Carbon dots; dye-doped silica nanoparticles |
The selection of appropriate reagents and materials must balance analytical performance with environmental considerations. For example, while traditional organic solvents like acetonitrile and methanol are effective for many extraction and separation processes, they pose environmental and safety concerns. Green alternatives include:
- Deep Eutectic Solvents (DES): Formed by mixing hydrogen bond donors and acceptors, these solvents are characterized by low toxicity, biodegradability, and often can be prepared from natural products [28]
- Bio-based Solvents: Derived from renewable biomass sources, these solvents offer reduced environmental impact throughout their lifecycle [29]
- Supercritical CO₂: Used particularly in extraction and chromatography, this solvent leaves no residual waste and is non-toxic [29]
Similarly, the move toward paper-based microfluidics represents a greener alternative to polymer-based devices, as paper is biodegradable, inexpensive, and requires minimal processing. These substrates can be functionalized with recognition elements for specific analytical applications while maintaining compatibility with smartphone-based detection.
Future Perspectives and Challenges
The continued advancement of smartphone-based PON analysis faces several technical and implementation challenges that represent opportunities for future research and development:
- Performance Validation: While numerous proof-of-concept studies have demonstrated the feasibility of smartphone-based detection, broader validation studies comparing these systems to established laboratory methods are needed, particularly for clinical applications where regulatory approval is required.
- Standardization and Reproducibility: Variations between smartphone models, camera specifications, and environmental conditions create challenges for achieving reproducible results. Developing calibration standards and normalization approaches will be essential for wider adoption.
- Multiplexing Capabilities: Most current systems focus on single-analyte detection, whereas many real-world applications require simultaneous measurement of multiple parameters. Developing multiplexed detection schemes within the constraints of smartphone optics represents an important frontier.
- Data Security and Privacy: As these systems increasingly incorporate patient or sensitive environmental data, ensuring secure data handling, transmission, and storage becomes critical.
- Integration with Artificial Intelligence: Machine learning and AI algorithms offer powerful approaches for enhancing image analysis, improving detection limits, and providing diagnostic decision support [19]. The convergence of smartphones with "smart assays and smart apps powered by machine learning and artificial intelligence holds immense promise for realizing a future for molecular analysis that is powerful, versatile, democratized" [19].
The environmental benefits of these systems could be further enhanced through:
- Life Cycle Assessment (LCA): Systematic evaluation of the environmental impact of PON systems from manufacturing through disposal would provide a more complete picture of their sustainability [29]
- Design for Disassembly and Recycling: Intentionally designing devices for easy separation of components and material recovery at end-of-life
- Renewable Energy Integration: Incorporating solar charging or other renewable energy sources for operation in off-grid settings
Despite these challenges, the trajectory of smartphone-based PON analysis points toward increasingly sophisticated, accessible, and sustainable analytical capabilities that have the potential to transform how chemical and biological measurements are performed across healthcare, environmental monitoring, and industrial applications.
Advanced Optical Methods and Their Biomedical Applications
Smartphone-Based Digital Image Colorimetry (SBDIA) for Pharmaceutical and Clinical Assays
Smartphone-based Lab-on-a-Chip (LoC) systems represent a transformative approach to molecular analysis, aiming to decentralize testing from central laboratories to the point-of-need. Within this framework, Smartphone-Based Digital Image Colorimetry (SBDIA) has emerged as a powerful and versatile optical detection method. It leverages the ubiquitous smartphone as a portable, cost-effective, and sophisticated analytical platform [19]. The core principle of SBDIA involves using a smartphone's camera to capture images of colored assay products, followed by the extraction of quantitative color intensity data using onboard apps or external software [30] [31]. This convergence of smartphones with optical assays democratizes analytical capabilities, making them accessible for use in resource-limited settings for pharmaceutical quality control and clinical diagnostics, thereby supporting a future of powerful, democratized molecular analysis [19].
Technical Foundations of SBDIA
The Smartphone as an Analytical Platform
The suitability of smartphones for colorimetric analysis stems from their highly integrated and advanced features. Modern smartphones are equipped with high-resolution cameras, built-in white LED lights for illumination, and substantial computational power for data processing [30]. Furthermore, features like wireless connectivity (Wi-Fi, Bluetooth) enable rapid transmission of results, while GPS can geo-tag measurements, which is valuable for environmental monitoring and supply chain tracking [19] [30]. This integration creates a complete analytical package that is both portable and user-friendly, eliminating the need for multiple bulky and expensive peripheral devices [19].
Core Colorimetric Detection Methods
SBDIA primarily relies on the measurement of color intensity resulting from a biochemical reaction. The process typically involves:
- Assay Reaction: A target analyte reacts with specific reagents to produce a colored compound.
- Image Acquisition: The smartphone camera captures an image of the colored solution, often under controlled lighting conditions (e.g., within a dark box) to minimize external interference [31].
- Color Quantification: The image is processed to extract color channel values. The most common color models are:
- RGB (Red, Green, Blue): The intensity of each primary color is represented by a value, typically from 0 to 255. The channel most responsive to the color change (e.g., the Blue channel for a yellow solution) is often selected for analysis [31].
- CMY (Cyan, Magenta, Yellow): Calculated as
CMY = 255 - RGB, these values are directly proportional to the concentration of a colored product, providing a more intuitive correlation with analyte concentration [31].
The extracted color values are then correlated with analyte concentration to generate a calibration curve and quantify unknown samples.
Experimental Protocols in SBDIA
Protocol 1: Determination of Peracetic Acid in Pharmaceutical Disinfectants
This method enables rapid quality control of disinfectant preparations at the point-of-use [32].
- Principle: Peracetic acid oxidizes iodide to iodine, which then reacts with N,N-diethyl-phenylenediamine to form a pink-magenta product. The intensity of this color is proportional to the peracetic acid concentration [32].
- Materials & Reagents:
- Peracetic acid standard solutions
- Potassium iodide (KI) solution
- N,N-diethyl-phenylenediamine solution
- 96-well microplate
- Smartphone with a custom-built app (e.g., "Modern Peracetic Acid Analysis") or generic color analysis app
- Procedure:
- Reaction: In a well of the microplate, mix the sample or standard peracetic acid solution with KI and N,N-diethyl-phenylenediamine.
- Incubation: Allow the reaction to proceed to develop the pink-magenta color.
- Imaging: Place the microplate on a uniform white background and capture an image of the entire plate using the smartphone camera, ensuring consistent lighting.
- Analysis: Use the smartphone app to analyze the relative green intensity of each well (the complementary color to magenta provides the highest sensitivity).
- Quantification: The concentration of peracetic acid in unknown samples is determined from a calibration curve of green intensity versus concentration (typical range: 0.15–3.0 µg/mL) [32].
The following diagram illustrates the workflow for this assay:
Protocol 2: Quantitative Determination of Uric Acid in Urine
This method provides a cost-effective alternative for clinical monitoring of uric acid levels, relevant for conditions like gout and renal disorders [31].
- Principle: Uric acid reduces phosphotungstic acid in an alkaline medium (sodium carbonate) to produce a characteristic blue color (tungsten blue) [31].
- Materials & Reagents:
- Uric acid standard solutions
- Phosphotungstate reagent
- Sodium carbonate (Na₂CO₃) solution (10%)
- Glass cuvettes or a multi-well plate
- Smartphone
- Computer with Image J software
- Procedure:
- Reaction: In a volumetric flask, mix the urine sample or standard with sodium carbonate solution. Let it stand for 10 minutes. Add phosphotungstate reagent, vortex mix, and dilute to volume.
- Imaging: Transfer the solutions to cuvettes. Place them in an imaging box with a white background to control lighting. Capture an image using a smartphone.
- Image Processing: Transfer the image to a computer. Open it in Image J. Crop the image to include all samples and convert it to an RGB stack.
- Color Quantification: Use Image J's "Plot Profile" function to measure the RGB gray values across each sample.
- Data Conversion: Convert the RGB values to CMY using the formula:
Cyan = 255 - R,Magenta = 255 - G,Yellow = 255 - B. The values from the channel most responsive to the blue color (e.g., Yellow) are used for quantification. - Quantification: Plot the CMY values against uric acid concentration to generate a calibration curve (typical range: 3–15 µg/mL) and determine the concentration in unknown samples [31].
The workflow for the uric acid assay is as follows:
Performance Data and Analytical Figures of Merit
The analytical performance of SBDIA methods is characterized by parameters such as linear range, limit of detection (LOD), limit of quantitation (LOQ), and precision. The following table summarizes these metrics for the featured assays and provides a comparison with a standard method.
Table 1: Analytical Performance of Representative SBDIA Methods
| Analyte | Matrix | Detection Method | Linear Range | LOD | LOQ | Comparison with Reference Method | Citation |
|---|---|---|---|---|---|---|---|
| Peracetic Acid | Pharmaceutical Disinfectant | Smartphone App (Green Intensity) | 0.15 - 3.0 µg/mL | 0.11 µg/mL | 0.34 µg/mL | No significant difference from traditional acid-base titration at 95% confidence level. | [32] |
| Uric Acid | Artificial Urine | Image J (CMY) | 3 - 15 µg/mL | Information Missing | Information Missing | Correlation coefficient nearly equivalent to UV/VIS spectrophotometry. | [31] |
| Uric Acid | Artificial Urine | Mobile App (B Channel) | 3 - 15 µg/mL | Information Missing | Information Missing | Lower correlation coefficient (0.97) compared to UV/VIS spectrophotometry. | [31] |
The Scientist's Toolkit: Essential Reagents and Materials
Successful implementation of SBDIA requires a set of key reagents and materials. The table below lists essential items and their functions in typical SBDIA workflows.
Table 2: Key Research Reagent Solutions for SBDIA
| Item | Function in SBDIA |
|---|---|
| N,N-diethyl-phenylenediamine | Chromogenic agent that is oxidized to form a pink-magenta dye, used in disinfectant testing. [32] |
| Potassium Iodide (KI) | Used as an intermediate in redox reactions; oxidized by peroxides to iodine, which then reacts with chromogens. [32] |
| Phosphotungstate Reagent | A phosphotungstic acid reagent used in clinical assays; reduced by analytes like uric acid to form a blue-colored product (tungsten blue). [31] |
| Sodium Carbonate (Na₂CO₃) | Provides an alkaline medium necessary for certain color development reactions, such as the reduction of phosphotungstate. [31] |
| 96-Well Microplate | A standard platform for running multiple assays in parallel, facilitating high-throughput analysis and consistent imaging. [32] |
| 3D-Printed Imaging Box / Cuvette Holder | Provides controlled, consistent lighting conditions during image capture, minimizing shadows and glare, which is critical for reproducibility. [30] |
Data Analysis and Validation
Advanced Analysis with Image J and Mobile Apps
While simple mobile apps can provide semi-quantitative analysis, advanced software like Image J offers superior quantitative capabilities. Image J allows for precise background subtraction, noise reduction, and intensity measurements across specific regions of interest, leading to more accurate and reliable data [31]. Studies have shown that analysis with Image J can yield correlation coefficients nearly equivalent to those from traditional UV/VIS spectrophotometry, outperforming results from some mobile apps which may be suitable only for qualitative or semi-quantitative analysis [31].
Method Validation and Greenness Assessment
Validating an SBDIA method against a standard reference method is crucial. For instance, the peracetic acid SBDIA method showed no significant statistical difference from classical acid-base titration [32]. Furthermore, the greenness of SBDIA methods can be evaluated using metrics like the Complementary Green Analytical Procedure Index and Analytical Greenness, which have demonstrated that SBDIA offers enhanced environmental friendliness and practical advantages over traditional methods due to its minimal reagent use and portable instrumentation [32].
Fluorescence detection has revolutionized biological and chemical analysis by providing exquisite sensitivity and specificity for detecting molecular events. This process is a three-stage cycle involving excitation, excited-state lifetime, and emission [33]. A photon of energy (hνEX) supplied by an external source is absorbed by a fluorophore, creating an excited electronic singlet state (S1') [33]. During the finite excited-state lifetime (typically 1-10 nanoseconds), the fluorophore undergoes conformational changes and interacts with its molecular environment, resulting in a relaxed singlet excited state (S1) from which fluorescence emission originates [33]. Finally, a photon of lower energy (hνEM) is emitted, returning the fluorophore to its ground state S0 [33].
The Stokes shift—the difference in energy or wavelength between excitation and emission photons—is fundamental to fluorescence sensitivity because it allows emission photons to be detected against a low background, isolated from excitation photons [33]. This physical process enables detection technologies ranging from ensemble measurements in microplate readers to the observation of individual biomolecules, with applications spanning clinical diagnostics, drug discovery, and fundamental biological research.
Core Physics of Fluorescence
Jablonski Diagram and Photophysics
The fluorescence process is comprehensively described by the Jablonski diagram, which illustrates the electronic states of a fluorophore and the transitions between them [33]. Upon light absorption, an electron is elevated to a higher energy state in a process characterized by a time scale of ∼10−15 s [34]. The excited electron then loses energy through vibrational relaxation over 10−14–10−11 s, followed by a transition back to the ground state with photon emission (10−9–10−7 s) [34]. This emitted photon has a longer wavelength than the incident light due to energy dissipation during the excited-state lifetime [33].
Key Fluorescence Parameters
Several spectroscopic parameters determine the utility of fluorescent probes for specific applications. The table below summarizes these critical properties and their significance in assay development.
Table 1: Key Fluorescence Properties and Their Significance
| Property | Definition | Significance in Detection |
|---|---|---|
| Extinction Coefficient | Capacity for light absorption at a specific wavelength | Determines brightness; fluorescence output is proportional to the product of extinction coefficient and quantum yield [33] |
| Quantum Yield (QY) | Number of fluorescence photons emitted per excitation photon absorbed | Directly impacts signal intensity; higher QY enables more sensitive detection [33] |
| Stokes Shift | Difference in energy/wavelength between excitation and emission photons | Enables separation of emission signal from excitation background; fundamental to sensitivity [33] |
| Photostability | Resistance to photochemical destruction during excitation | Critical for prolonged imaging and single-molecule tracking; limits observation time [34] |
| Fluorescence Lifetime | Average time the molecule spends in excited state before emission | Enables fluorescence lifetime imaging (FLIM) and discrimination of environmental changes [35] |
The entire fluorescence process is cyclical, and unless the fluorophore is irreversibly destroyed (photobleaching), the same fluorophore can be repeatedly excited, generating many thousands of detectable photons—a fundamental aspect enabling the high sensitivity of fluorescence detection techniques [33].
Established High-Sensitivity Fluorescence Technologies
Fluorescence Immunoassays
Fluorescence detection has significantly enhanced conventional immunoassay platforms. Traditional enzyme-linked immunosorbent assay (ELISA) has been adapted to enzyme-linked fluorescence assay (ELFA) by replacing colorimetric substrates with fluorescent counterparts like 4-methylumbelliferyl phosphate [36]. This modification provides substantial sensitivity advantages, allowing assays to be conducted with less antigen or shorter substrate incubation times (5 minutes for ELFA versus 30 minutes for ELISA for rubella antibody detection) [36].
Recent comparative studies demonstrate the performance advantages of fluorescence-based immunoassays. In dengue virus detection, fluorescent immunoassay (FIA) showed slightly superior performance to immunochromatography (IC), with sensitivity of 79.11% versus 76.58% for NS1 antigen detection, while maintaining equal specificity at 92.28% [37]. The FIA platform also demonstrated higher positive predictive value (86.81% vs. 86.43%), negative predictive value (87.31% vs. 85.98%), and overall agreement (87.13% vs. 86.14%) compared to conventional immunochromatography [37].
Fluorescence Detection Instrumentation
Fluorescence detection systems share four essential elements: (1) an excitation light source, (2) a fluorophore, (3) wavelength filters to isolate emission photons from excitation photons, and (4) a detector that registers emission photons [33]. These components are configured differently across specialized instruments:
- Spectrofluorometers and microplate readers measure average properties of bulk samples (µL to mL) [33]
- Fluorescence microscopes resolve fluorescence as a function of spatial coordinates in microscopic objects [33]
- Fluorescence scanners resolve fluorescence in macroscopic objects like gels and blots [33]
- Flow cytometers measure fluorescence per cell in a flowing stream [33]
Each instrument type produces different measurement artifacts and imposes different demands on fluorescent probes. For example, photobleaching is often significant in fluorescence microscopy but less problematic in flow cytometry due to short dwell times in the excitation beam [33].
Advanced Single-Molecule Fluorescence Techniques
Principles and Methodologies
Single-molecule fluorescence microscopy (SMFM) enables the investigation of biological structure and function at the ultimate sensitivity limit—observing individual molecules [38]. This approach reveals information about molecular behavior that would otherwise be hidden in ensemble averages, where subpopulations and rare events are obscured [34]. SMFM has uncovered fundamental processes including protein folding, DNA replication, bacterial flagellar motor rotation, and viral infection mechanisms [34].
SMFM requires fluorophores that are exceptionally bright, photostable, and small to avoid disrupting biological activity [38]. The most popular fluorophores include organic dyes (FITC, TRITC), fluorescent proteins (GFP, YFP), and quantum dots [38]. Advanced labeling strategies have been developed for single-molecule work, as illustrated in the methodology diagram below.
Key Applications and Workflows
Single-molecule techniques are particularly valuable for studying cellular heterogeneity and molecular dynamics. While traditional experimental investigations are performed on population "ensemble average" levels, this approach risks losing valuable information concerning biologically relevant heterogeneity, such as drug-resistant bacteria or cancer cells in a general cellular population [34]. SMFM enables researchers to identify and investigate molecular subpopulations within cells, studying not only cellular responses but also precise underlying molecular mechanisms [34].
A representative single-molecule tracking workflow for studying protein dynamics in live cells involves several critical steps, from sample preparation through data analysis, with particular attention to minimizing background noise and optimizing signal-to-noise ratio for reliable single-particle tracking.
Table 2: Essential Research Reagents for Single-Molecule Fluorescence Studies
| Reagent Category | Specific Examples | Function and Application |
|---|---|---|
| Fluorescent Proteins | GFP, YFP, mEos, Dendra, mCherry | Genetically-encoded tags for protein labeling and localization in live cells [34] [39] |
| Organic Dyes | FITC, TRITC | Bright, photostable small molecules for in vitro and fixed cell labeling [38] |
| Labeling Methods | Antibodies, Biotin-Streptavidin, Epitope tags | Covalent and non-covalent attachment strategies for specific molecular targeting [38] |
| Photoactivatable Proteins | PA-GFP, Dendra2, mMaple | Enable super-resolution techniques (PALM/STORM) through controlled activation [34] |
| FRET Biosensors | CFP-YFP pairs | Detect molecular interactions and conformational changes via energy transfer [39] |
Smartphone-Based Fluorescence Detection for Lab-on-Chip Applications
Technical Foundations and Advantages
Smartphones have emerged as powerful platforms for portable fluorescence detection in lab-on-chip (LOC) applications, leveraging their ubiquitous penetration, integrated features, and advanced cameras [19]. With approximately 54% of the world's population owning a smartphone and mobile networks available to 95%, this technology offers unprecedented accessibility for point-of-care diagnostics [19]. The economy of scale in smartphone manufacturing ($500 billion USD market) enables costs far lower than bespoke scientific instruments, making advanced detection technology financially accessible [19].
Modern smartphones integrate numerous components directly useful for fluorescence measurements: high-resolution cameras with sensitive sensors, intense LED flashes for excitation, powerful processors for data analysis, wireless connectivity for data transmission, and touchscreen interfaces for user interaction [19]. These features create a complete technological package that minimizes the size, weight, and complexity of portable fluorescence detection systems compared to microcontroller unit (MCU) or single-board computer (SBC) alternatives [19].
Implementation Approaches
Smartphone-based fluorescence detection systems typically interface with custom-designed components that accommodate specific assay requirements. These include:
- 3D-printed adapters for precise alignment of samples and optical components
- External lenses for magnification and improved light collection
- Custom filters to separate excitation and emission wavelengths
- Microfluidic chips for sample handling and processing
- Portable light sources for uniform excitation when higher intensity is required
The high sensitivity of smartphone cameras continues to improve, with recent models featuring larger sensors, better low-light performance, and advanced computational photography algorithms that can be leveraged for quantitative fluorescence measurements [19]. This capabilities enable smartphone-based systems to approach the performance of conventional laboratory instruments for many applications, particularly in point-of-care settings where rapid results are critical.
Comparative Performance Analysis
Sensitivity Across Platforms
The evolution of fluorescence detection technologies has progressively enhanced sensitivity, enabling applications from ensemble measurements to single-molecule detection. The table below compares the key characteristics and performance metrics across this sensitivity spectrum.
Table 3: Performance Comparison of Fluorescence Detection Platforms
| Technology | Detection Limit | Key Applications | Advantages | Limitations |
|---|---|---|---|---|
| Fluorescence ELISA | ~pM concentrations | Clinical diagnostics, pathogen detection [36] [37] | Quantitative, well-established, high throughput | Requires multiple washing steps, moderate sensitivity |
| Fluorescent Immunoassay (FIA) | Enhanced sensitivity over colorimetric | Rapid diagnostic testing (e.g., dengue NS1 detection) [37] | Faster than ELISA (5-15 min vs 30 min), higher sensitivity than immunochromatography [36] [37] | Limited multiplexing, requires reader instrumentation |
| Confocal Microscopy | Single molecules in small volumes | Cellular imaging, fluorescence correlation spectroscopy [35] | Optical sectioning, reduced background, high spatial resolution | Complex instrumentation, limited field of view |
| Single-Molecule Microscopy | Individual biomolecules | Molecular tracking, super-resolution imaging, heterogeneity studies [34] [38] | Ultimate sensitivity, reveals heterogeneity, molecular counting | Specialized fluorophores required, technical complexity |
| Smartphone Detection | Variable (platform-dependent) | Point-of-care testing, field deployment [19] | Portability, accessibility, cost-effectiveness, connectivity | Limited sensitivity vs. dedicated instruments |
Technical Considerations for Implementation
Implementing high-sensitivity fluorescence detection requires careful consideration of multiple technical factors. Fluorescence intensity is quantitatively dependent on the molar extinction coefficient, optical path length, solute concentration, fluorescence quantum yield, excitation source intensity, and fluorescence collection efficiency of the instrument [33]. In dilute solutions or suspensions, fluorescence intensity is linearly proportional to these parameters, but when sample absorbance exceeds approximately 0.05 in a 1 cm pathlength, the relationship becomes nonlinear due to artifacts like self-absorption and the inner-filter effect [33].
For quantitative applications, reference standards are essential for calibrating measurements made at different times or using different instrument configurations [33]. High-precision fluorescent microsphere standards are available for fluorescence microscopy and flow cytometry, while ready-made fluorescent standard solutions facilitate spectrofluorometer calibration [33]. These standards ensure measurement consistency and enable reliable comparison of results across platforms and laboratories.
Future Perspectives
The convergence of smartphones with smart assays and smart apps powered by machine learning and artificial intelligence holds immense promise for realizing a future for molecular analysis that is powerful, versatile, and democratized [19]. Ongoing development of brighter, more photostable fluorophores—particularly in the far-red and near-infrared regions—will further enhance sensitivity while reducing background autofluorescence in biological samples [39] [40].
Advanced fluorescence fluctuation techniques like fluorescence correlation spectroscopy (FCS) and photon counting histogram (PCH) analysis are pushing detection limits in high-throughput screening applications, enabling researchers to investigate biomolecular interactions at previously inaccessible resolution [41] [35]. These developments, combined with miniaturized detection platforms, promise to transform molecular analysis from a specialized laboratory technique to a widely accessible tool for health monitoring, environmental sensing, and fundamental biological discovery.
Label-free detection techniques represent a cornerstone of modern bioanalysis, enabling researchers to study biomolecular interactions in their native state without the need for fluorescent, radioactive, or other modifying labels. These methods monitor interactions in real-time by measuring inherent physicochemical properties of molecules, such as mass, refractive index, or dielectric properties [42] [43]. This approach preserves natural molecular behavior and provides direct access to binding kinetics and affinity constants, overcoming the significant limitation of label-based techniques where the labeling process can alter molecular structure and function [44] [45]. The elimination of labeling steps simplifies assay protocols, reduces preparation time and costs, and enables the study of molecular systems where labeling is impractical or would interfere with binding sites [42] [43].
The field has evolved substantially from initial surface plasmon resonance (SPR) systems to encompass a diverse range of technological approaches including interferometry, grating-coupled interferometry (GCI), microcantilevers, and nanoplasmonic sensing [45] [46]. Recent advancements have pushed detection limits to unprecedented sensitivities, achieving single-molecule detection in some configurations [44] [46]. These technological improvements, combined with the inherent advantages of observing unmodified biomolecules, have established label-free techniques as indispensable tools across fundamental biological research, drug discovery, diagnostic development, and increasingly, point-of-care testing platforms [45] [46].
Fundamental Principles of Optical Label-Free Detection
Optical label-free detection techniques fundamentally rely on monitoring changes in the local refractive index that occur when biomolecules interact with a functionalized sensor surface [46]. When a target analyte binds to its immobilized recognition partner, the accumulation of biomolecular mass alters the optical properties at the surface-solution interface. This phenomenon forms the basis for detecting binding events without labels by measuring the resulting perturbation of incident light [44]. The detection principle capitalizes on the contrast between the refractive index of biomolecules (typically n ~ 1.59 for proteins) and their surrounding aqueous medium (n ~ 1.33) [44].
For deeply sub-diffractional particles like proteins, the light-matter interaction is governed by non-resonant processes characterized by the molecule's polarizability, which quantifies its ability to deform its electron cloud in response to an incident electromagnetic field [44]. The direct manifestation of polarizability is scattering and absorption, though absorption is typically negligible for most biomolecules in the visible spectrum except for naturally absorbing species like hemoglobin or GFP [44]. For small, non-absorbing particles up to a tenth of the wavelength of light, this interaction is governed by elastic Rayleigh scattering, where the scattering cross-section (a measure of scattered light intensity) is proportional to the square of the polarizability and scales linearly with molecular volume [44]. The inherently low refractive index contrast between biomolecules and aqueous environments results in weak scattering signals that diminish dramatically with size, scaling with the sixth power of the particle diameter [44]. This relationship presents a significant challenge for conventional optical detection methods, necessitating sophisticated signal enhancement strategies for detecting small biomolecules at low concentrations.
Signal Enhancement Strategies
To overcome the fundamental limitation of weak scattering signals from biomolecules, several advanced signal enhancement strategies have been developed:
Interference Enhancement: Techniques like Interference Scattering Microscopy (iSCAT) leverage wave interference between light scattered by a biomolecule and a coherent reference wave to enhance weak signals [44]. The total detected intensity (It) follows the principle: It = |Er|² + |Es|² + 2|Er||Es|cosϕ, where Er is the reference wave field, Es is the scattered wave field, and ϕ is the phase difference between them. For subwavelength particles, the interference term becomes the main contributor to the detected signal after background subtraction [44].
Plasmonic Enhancement: Metallic nanostructures support localized surface plasmon resonances (LSPR) that generate enhanced electromagnetic fields at their surfaces, significantly amplifying signals from nearby biomolecules [47] [48]. The antenna effect of plasmonic particles can confine and enhance light-matter interactions, enabling detection of minute quantities of biomaterials [44].
Resonance Enhancement: Optical resonance phenomena in structures such as Fabry-Pérot cavities and whispering gallery mode resonators create standing waves that enhance the interaction between light and molecules, improving detection sensitivity [44].
Nanoscale Field Confinement: High-field enhancement in nanoscale apertures in metal films can concentrate electromagnetic energy into volumes much smaller than the wavelength of light, dramatically increasing detection sensitivity for single molecules [44].
Major Label-Free Detection Technologies
Surface Plasmon Resonance (SPR) and Localized SPR
Surface Plasmon Resonance (SPR) stands as the most established and widely utilized label-free technology for biomolecular interaction analysis [42]. In SPR, a thin gold film is excited by incident light under specific conditions, generating electromagnetic waves (surface plasmons) that propagate along the metal surface [42]. When biomolecules bind to the functionalized surface, the local refractive index changes, altering the resonance conditions and causing a detectable shift in the resonance angle or wavelength [44]. SPR provides real-time monitoring of binding events, enabling determination of association (ka) and dissociation (kd) rate constants, and calculation of the equilibrium dissociation constant (K_D) [42] [45].
Localized Surface Plasmon Resonance (LSPR) employs noble metal nanoparticles rather than continuous metal films [47]. When incident light matches the natural frequency of surface electrons in these nanoparticles, it generates LSPR with strongly enhanced local electromagnetic fields [47]. LSPR offers several advantages over conventional SPR, including greater field enhancements, miniaturization potential, and simplified optical setups [47]. The resonance conditions in LSPR are highly sensitive to parameters such as particle size, shape, interparticle spacing, and changes in the local refractive index caused by molecular binding events [47].
Table 1: Comparison of Major Label-Free Detection Technologies
| Technology | Principle | Applications | Advantages | Limitations |
|---|---|---|---|---|
| SPR | Measures refractive index changes via surface plasmon waves on thin metal films | Biomolecular interaction analysis, kinetic studies, drug discovery | Real-time measurements, sensitive to conformational changes, quantitative | Restricted to gold/silver surfaces, requires sophisticated instrumentation |
| LSPR | Utilizes localized plasmons on nanoparticles for refractive index sensing | Protein-protein interactions, small molecule screening, diagnostic assays | Miniaturization capability, simpler optics, high field enhancement | Smaller detection volume, more complex surface chemistry |
| Interferometry | Measures interference patterns between reference and sample beams | Single-molecule detection, mass quantification, interaction kinetics | High sensitivity, quantitative mass measurement, single-molecule capability | Sensitive to environmental noise, complex data interpretation |
| Grating-Coupled Interferometry (GCI) | Uses diffraction gratings to create interference patterns | Drug discovery, immunogenicity studies, vaccine development | High sensitivity, suitable for high-throughput screening | Specialized sensor chips required |
| SEIRA | Enhances molecular vibrational signals via plasmonic nanostructures | Chemical-specific detection, dynamic monitoring of biomolecules | Molecular structural information, fingerprint identification | Limited to IR-active vibrations, complex nanostructure fabrication |
Interferometric Methods
Interferometric detection techniques leverage the wave nature of light to achieve exceptional sensitivity in biomolecular detection. Interference Scattering Microscopy (iSCAT) has emerged as a leading interferometric method, capable of detecting single proteins in the tens of kilodalton range [44]. The contrast in iSCAT scales linearly with protein mass, functioning as an optical analog of mass spectrometry that enables precise mass profiling and real-time tracking of molecular transport [44].
Various implementations of interference microscopy exist with different illumination and detection schemes, including reflection-based configurations where most incident light passes through the substrate while a small fraction reflects and interferes with back-scattered light from biomolecules [44]. Transmission-based interferometry methods like Coherent Brightfield Imaging (COBRI) rely on interference between forward-scattered light and an unscattered transmitted beam, reducing the impact of reflections from internal interfaces and making them advantageous for tracking objects in complex environments like whole cells [44].
A novel technique called Nanofluidic Scattering Microscopy (NSM) addresses some limitations of traditional interference microscopy by employing nanochannels as containers for freely diffusing molecules [44]. In NSM, the reference wave originates from light scattered by the nanochannel walls, ensuring minimal axial displacement of molecules and preventing full-phase oscillations, resulting in a stable signal throughout imaging while allowing diffusivity measurements alongside molecular mass quantification [44].
Surface-Enhanced Infrared Absorption (SEIRA) Spectroscopy
Surface-Enhanced Infrared Absorption (SEIRA) spectroscopy represents an advanced label-free technique that combines the molecular specificity of infrared spectroscopy with the enhancement capabilities of plasmonic metasurfaces [48]. SEIRA addresses the fundamental challenge of weak IR light absorption by molecules—typically with absorption coefficients around 10³ M⁻¹cm⁻¹, significantly lower than electronic transitions in UV-visible ranges (approximately 10⁶ M⁻¹cm⁻¹) [48].
Recent innovations in SEIRA include the development of dual-band plasmonic metasurfaces based on surface lattice resonances that simultaneously enhance molecular vibrations at multiple frequencies [48]. These platforms can be engineered to match specific molecular vibrations, such as methyl and amide bands in proteins, enabling comprehensive analysis of biomolecular interactions [48]. The electric fields on these mixed arrays can be strongly confined to approximately 100 nm, enabling high SEIRA performance and approaching single-molecule detection capabilities [48].
SEIRA has been successfully applied to monitor biomolecular interactions in real-time, such as between protein A and immunoglobulin G (IgG), providing remarkable changes in the intensity and vibrational features of SEIRA absorbance [48]. These dynamic SEIRA measurements use molecular vibrations as self-biomarkers, identifying kinetic parameters for important affinity metrics of biomolecular interactions, including association (ka) and dissociation (kd) rate constants [48].
Diagram 1: SEIRA Biosensing Workflow. This flowchart illustrates the operational process of Surface-Enhanced Infrared Absorption (SEIRA) spectroscopy for biomolecular interaction analysis, from illumination to affinity constant determination.
Integration with Smartphone-Based Lab-on-Chip Platforms
The convergence of label-free detection technologies with smartphone-based platforms represents a cutting-edge development in point-of-care testing and decentralized diagnostics [19] [49]. Smartphones offer an ideal foundation for portable analytical devices due to their integrated technological package, including high-resolution cameras, powerful processors, connectivity options, and user-friendly interfaces [19]. The global ubiquity of smartphones—with approximately 54% of the world's population owning one—creates unprecedented opportunities for democratizing sophisticated biomolecular analysis [19].
Smartphone cameras serve as sophisticated optical detectors that can be leveraged for various label-free detection modalities [19]. Modern smartphone cameras incorporate advanced features including autofocus systems, image stabilization, programmable exposure, and ISO sensitivity settings that make them suitable for scientific applications [19]. When combined with specialized attachments and microfluidic chips, smartphones can function as portable laboratories capable of performing sophisticated analyses outside traditional laboratory settings [19] [49].
The motivation for adopting smartphones as platforms for molecular analysis stems from several key advantages: their global penetration across diverse populations, the economy of scale that reduces costs, and their integrated package of sensors, processors, and communication capabilities [19]. These features collectively address the critical need for LOC technologies that offer rapid quantitative analysis, straightforward operation, and democratization of access without substantial financial barriers [19].
Implementation Approaches
Smartphone-based label-free detection systems typically follow one of several implementation approaches:
Camera-Based Detection: Utilizing the smartphone's built-in camera as the primary detector for optical signals, often coupled with specialized attachments that interface with microfluidic chips or sensor surfaces [19] [49]. The camera can capture changes in interference patterns, refractive index variations, or light scattering that indicate molecular binding events.
Wired Peripherals: Connecting specialized sensor modules to smartphones through USB interfaces or audio jacks, enabling electrochemical detection or providing power to external components [49]. This approach maintains the smartphone's portability while expanding its analytical capabilities.
Wireless Connectivity: Leveraging Bluetooth, Wi-Fi, or NFC for data transfer between external sensor modules and smartphones, eliminating the need for physical connections and enhancing user convenience [49]. This approach is particularly valuable for wearable sensors or continuous monitoring applications.
Table 2: Smartphone-Based Detection Modalities for Label-Free Analysis
| Detection Modality | Measurement Principle | Compatible Label-Free Techniques | Smartphone Components Utilized |
|---|---|---|---|
| Imaging-Based | Capture of optical signals: interference patterns, scattering, refractive index changes | Interferometry, iSCAT, NSM, LSPR imaging | Built-in camera, flash, processor |
| Spectroscopic | Spectral analysis of reflected or transmitted light | SPR, LSPR, SEIRA, reflectance spectroscopy | Camera with diffraction gratings, light source |
| Electrochemical | Measurement of electrical impedance changes | Impedance spectroscopy, field-effect sensing | Audio jack for data acquisition, USB for power |
| Connected Modules | External sensors with data transmission | GCI, waveguide-based sensors, calorimetry | Bluetooth, Wi-Fi, USB for communication |
Experimental Protocols and Methodologies
Surface Functionalization for Label-Free Biosensing
Proper surface functionalization is critical for successful label-free biomolecular interaction analysis. The following protocol outlines a standard approach for preparing biosensor surfaces:
Surface Cleaning: Thoroughly clean the sensor surface (typically gold for SPR) using oxygen plasma treatment or piranha solution (3:1 concentrated H₂SO₄:30% H₂O₂) to remove organic contaminants. Caution: Piranha solution is highly corrosive and must be handled with appropriate safety measures.
Self-Assembled Monolayer (SAM) Formation: Immerse the clean sensor surface in a 1 mM solution of alkanethiols (e.g., 16-mercaptohexadecanoic acid) in ethanol for 12-24 hours to form a well-ordered SAM. The SAM provides functional groups for subsequent biomolecule immobilization and helps minimize nonspecific binding.
Activation of Carboxyl Groups: Treat the SAM-coated surface with a mixture of 0.4 M EDC (1-ethyl-3-(3-dimethylaminopropyl)carbodiimide) and 0.1 M NHS (N-hydroxysuccinimide) in water for 30 minutes to activate carboxyl groups, forming amine-reactive NHS esters.
Ligand Immobilization: Incubate the activated surface with the ligand solution (typically at 10-100 μg/mL in 10 mM acetate buffer, pH 5.0) for 30-60 minutes. The optimal pH should be 1-1.5 units below the ligand's isoelectric point to ensure positive charge and enhanced binding to the negatively charged surface.
Surface Blocking: Treat the functionalized surface with 1 M ethanolamine hydrochloride (pH 8.5) for 15 minutes to deactivate remaining activated groups and reduce nonspecific binding. Alternatively, bovine serum albumin (BSA) solutions (1% w/v) can be used as blocking agents.
Buffer Conditioning: Equilibrate the prepared biosensor with running buffer (typically PBS or HEPES-buffered saline) until a stable baseline is achieved before introducing analytes.
This protocol creates a well-defined biosensor surface capable of specifically capturing target analytes while minimizing nonspecific interactions, which is essential for obtaining reliable kinetic data [42] [48].
Kinetic Analysis of Biomolecular Interactions
Label-free technologies uniquely enable real-time monitoring of biomolecular interactions, providing direct access to kinetic parameters. The standard methodology for kinetic analysis involves:
Baseline Establishment: Flow running buffer across the functionalized sensor surface until a stable baseline is achieved, typically requiring 5-10 minutes of stabilization.
Association Phase Monitoring: Introduce the analyte at various concentrations (typically spanning a 10-100 fold range around the expected K_D) and monitor the binding response in real-time. The association phase should continue until binding approaches saturation or a predefined maximum time.
Dissociation Phase Monitoring: Replace the analyte solution with running buffer and monitor the decrease in response as complexes dissociate. The dissociation phase should continue until sufficient data is collected for accurate k_d determination, or until the next injection cycle.
Surface Regeneration: Apply a regeneration solution (typically mild acid or base) to remove bound analyte without damaging the immobilized ligand. Glycine-HCl (pH 2.0-3.0) or NaOH (10-100 mM) are commonly used regeneration solutions.
Data Analysis: Fit the resulting sensorgram data to appropriate binding models. For 1:1 interactions, the data is typically fit to the following equations:
During association: dR/dt = ka × C × (Rmax - R) - k_d × R
During dissociation: dR/dt = -k_d × R
Where R is the response, C is the analyte concentration, Rmax is the maximum binding capacity, ka is the association rate constant, and k_d is the dissociation rate constant.
Affinity Calculation: Determine the equilibrium dissociation constant (KD) from the ratio of the rate constants: KD = kd/ka [45] [48].
This methodology provides a comprehensive characterization of biomolecular interactions, revealing not just binding affinity but also the kinetics of complex formation and dissociation, which often correlates with biological efficacy.
Diagram 2: Kinetic Analysis Workflow. This flowchart outlines the standard process for determining kinetic parameters of biomolecular interactions using label-free biosensors, from baseline establishment to parameter calculation.
The Scientist's Toolkit: Essential Research Reagents and Materials
Successful implementation of label-free detection techniques requires specific reagents and materials optimized for each technology platform. The following table details essential components for establishing label-free biomolecular interaction analysis:
Table 3: Essential Research Reagents and Materials for Label-Free Biosensing
| Reagent/Material | Function | Application Notes | Compatible Techniques |
|---|---|---|---|
| Gold Sensor Chips | Provides surface for biomolecule immobilization and plasmon excitation | Typically coated with 47-50 nm gold film on glass with 1-2 nm chromium or titanium adhesion layer | SPR, LSPR, SEIRA |
| Alkanethiols | Form self-assembled monolayers (SAMs) for functionalizing gold surfaces | 16-Mercaptohexadecanoic acid commonly used for carboxyl-terminated surfaces | SPR, LSPR, SEIRA |
| Coupling Reagents (EDC/NHS) | Activate carboxyl groups for covalent immobilization of ligands | EDC concentration typically 0.4 M with 0.1 M NHS in water; 30-min activation | All covalent immobilization |
| Ethanolamine HCl | Blocks remaining activated groups after ligand immobilization | 1 M solution, pH 8.5; 15-min incubation | All techniques requiring surface blocking |
| CM-Dextran | Creates hydrophilic hydrogel matrix on sensor surfaces | Enhances immobilization capacity; reduces nonspecific binding | SPR, GCI |
| Protein A/G | Oriented immobilization of antibodies through Fc region | Improves antigen binding capacity; maintains antibody activity | SPR, BLI, GCI |
| Regeneration Solutions | Removes bound analyte without damaging immobilized ligand | Glycine-HCl (pH 2.0-3.0) or NaOH (10-100 mM); optimization required | All techniques requiring surface regeneration |
| HBS-EP Buffer | Standard running buffer for biomolecular interaction studies | 10 mM HEPES, 150 mM NaCl, 3 mM EDTA, 0.05% surfactant P20, pH 7.4 | SPR, GCI, interferometry |
| Plasmonic Nanoparticles | Enhance sensitivity through signal amplification | Gold nanospheres (20-100 nm), nanorods, nanostars; functionalized with recognition elements | LSPR, nanoparticle-enhanced SPR |
Applications in Biomolecular Analysis and Future Perspectives
Label-free detection techniques have found diverse applications across multiple domains of biological research and diagnostic development. In drug discovery, these methods enable characterization of candidate compound interactions with therapeutic targets, providing critical kinetic and affinity data that inform structure-activity relationships [45]. Pharmaceutical researchers utilize label-free technologies to study small molecule-protein interactions, antibody-antigen binding, and receptor-ligand engagements with unprecedented precision [45] [46]. The real-time monitoring capability allows researchers to distinguish promising drug candidates based not only on binding affinity but also on complex kinetic profiles that may correlate with in vivo efficacy [45].
In diagnostic applications, label-free biosensors are advancing toward clinical implementation, particularly for detection of disease biomarkers in complex biological fluids [46]. The COVID-19 pandemic accelerated development of label-free sensors for viral detection, with demonstrations of SARS-CoV-2 spike protein detection at concentrations as low as 430 fg/mL in saliva using smartphone-compatible platforms [46]. Similar approaches are being applied to cancer biomarker detection, cardiac marker analysis, and inflammatory indicator monitoring [47] [49]. The elimination of labeling steps simplifies assay protocols, reducing time-to-result and making these platforms particularly attractive for point-of-care testing scenarios [49] [46].
Emerging applications include single-molecule detection capabilities that push the boundaries of analytical sensitivity [44]. Advanced interferometric methods like iSCAT can now detect single proteins with masses in the tens of kilodalton range, enabling researchers to observe heterogeneities and transient states invisible to conventional ensemble measurements [44]. Similarly, refined SEIRA platforms with dual-band plasmonic metasurfaces can monitor biomolecular interactions while simultaneously tracking multiple vibrational modes, providing comprehensive insight into binding-induced structural changes [48]. These capabilities open new possibilities for studying low-abundance biomarkers, rare cellular events, and fundamental molecular processes at previously inaccessible resolution levels.
The future evolution of label-free detection technologies will likely focus on several key areas: further sensitivity enhancements to expand single-molecule applications, increased multiplexing capabilities for parallel analysis of multiple biomarkers, improved integration with portable platforms like smartphones for decentralized testing, and enhanced data analysis methodologies incorporating machine learning for extracting subtle binding signatures from complex datasets [19] [47] [46]. As these technologies mature and become more accessible, they are poised to transform biomolecular interaction analysis across basic research, drug development, and clinical diagnostics.
The field of optical detection is undergoing a transformative shift with the integration of smartphone-based platforms into laboratory-grade analytical methods. This evolution is guided by the principles of Green Analytical Chemistry (GAC), which advocates for the development of portable, cost-effective, and in-situ analysis techniques [50]. Smartphones, with their advanced image sensors, significant processing power, and inherent connectivity, offer an unparalleled platform for decentralizing chemical and biological analysis [50] [51]. This whitepaper examines two groundbreaking capabilities emerging from this convergence: portable super-resolution microscopy and digital bioassays. Both technologies leverage the smartphone's optical hardware and computational resources to achieve performance levels once restricted to expensive, centralized laboratory equipment. We detail the technical principles, experimental protocols, and practical implementations of these methods, framing them within the broader context of optical detection in smartphone-based lab-on-chip (LoC) research for drug development and diagnostic applications.
Smartphone-Based Portable Super-Resolution Microscopy
Principle and Microscope Design
Super-resolution microscopy techniques bypass the diffraction limit of light, allowing for optical resolution at the nanoscale [52]. A landmark advancement in this field is the development of a low-cost, portable smartphone-based fluorescence microscope capable of direct single-molecule detection without signal amplification [53]. This capability is foundational to techniques like Single-Molecule Localization Microscopy (SMLM), including DNA-PAINT, which achieve super-resolution by temporally separating the emission of individual fluorophores to precisely determine their positions [53] [52].
The smartphone microscope is a standalone unit (Figure 1) weighing approximately 1.2 kg with dimensions of 11 × 22 × 12 cm. Its design prioritizes sensitivity, portability, and affordability, with a total component cost under €350 [53]. The optical path is engineered to minimize background signal:
- Excitation: A laser beam is focused and directed through a half-ball lens to achieve Total Internal Reflection (TIR) or Highly Inclined and Laminated Optical sheet (HILO) illumination. This configuration minimizes background fluorescence from the sample volume [53].
- Emission: The fluorescence light from the sample is collected by a low numerical aperture (NA) air objective, filtered by an emission filter to block scattered laser light, and focused onto the smartphone's CMOS sensor by the smartphone's own camera lens, which acts as a tube lens [53].
- Modularity: The design features interchangeable stages for the laser, objective, and sample, making it compatible with various smartphone models and allowing for easy swapping of excitation wavelengths [53].
Figure 1. Optical pathway of the smartphone-based super-resolution microscope. The pathway illustrates laser excitation via total internal reflection (TIR) and subsequent fluorescence collection through the objective and emission filter onto the smartphone sensor.
Performance Metrics and Experimental Validation
The performance of this portable microscope was rigorously validated through single-molecule fluorescence experiments and super-resolution imaging.
- Single-Molecule Detection: Using DNA origami structures labeled with a single ATTO 647N dye molecule, the microscope demonstrated the ability to detect single-molecule emission with a signal-to-noise ratio (SNR) of approximately 3.3. The characteristic single-step photobleaching of individual dye molecules was clearly observed [53].
- Super-Resolution Imaging: By implementing DNA-PAINT, the microscope achieved super-resolution imaging of DNA origami structures and cellular microtubule networks. The system demonstrated a localization precision of 84 nm, resulting in a 6.6-fold enhancement in spatial resolution compared to the conventional diffraction limit [53].
- Versatility: The microscope's performance was benchmarked using three different commercially available smartphone models from Apple, Samsung, and Huawei, confirming its broad adaptability [53].
Table 1. Performance metrics of the smartphone-based super-resolution microscope.
| Parameter | Performance Value | Experimental Context |
|---|---|---|
| Single-Molecule SNR | 3.3 | Detection of ATTO 647N on DNA origami [53] |
| Localization Precision | 84 nm | DNA-PAINT imaging [53] |
| Resolution Enhancement | 6.6-fold | Compared to diffraction limit [53] |
| Cost | < €350 | Component cost [53] |
| Weight & Dimensions | 1.2 kg, 11x22x12 cm | Portable, standalone unit [53] |
Digital Bioassays Enabled by Smartphone Detection
Fundamental Principles and Advantages
Digital bioassays represent a paradigm shift from traditional analogue assays by isolating and detecting individual target molecules, transforming analog concentrations into digital counts [54]. This approach, when integrated with smartphone detection, offers several transformative advantages for analytical science and diagnostics [54]:
- High Sensitivity and Low Detection Limits: By dividing a sample into thousands of micro-compartments, digital assays can isolate single molecules, pushing the limit of detection to the attomolar range or lower. This allows for the detection of rare biomarkers, such as circulating tumor DNA for early cancer diagnosis [54].
- Quantitative Precision: Digital assays count discrete events, providing absolute quantification and highly reproducible data. This precision is vital for tracking viral load in outbreaks or making critical diagnostic decisions [54].
- Small Sample Volumes: Leveraging microfluidic technologies, these assays can perform comprehensive analyses on minute sample volumes, which is crucial in paediatrics or research involving rare biological samples [54].
- Automation and Scalability: The processes are amenable to automation, reducing manual labour and errors. Their inherent scalability supports high-throughput screening for drug discovery [54].
Smartphone Implementation of Digital Assays
The smartphone-based microscope previously described directly enables digital bioassays. In one demonstration, the system was used to implement a single-molecule bioassay for the detection of Ebola RNA fragments via DNA-PAINT, highlighting its potential for point-of-care (POC) diagnostics [53]. Beyond fluorescence microscopy, smartphones have been integrated with other detection modalities to create portable digital assay platforms:
- Electrochemical Immunosensors: A smartphone-based, label-free electrochemical immunosensor was developed for the detection of carcinoembryonic antigen (CEA), a cancer biomarker. The sensor uses a layer-by-layer assembly of nanostructured materials on a screen-printed electrode to amplify the signal. The smartphone interface allows for POC detection with a linear range of 0.1–5.0 ng mL⁻¹ and a low detection limit of 0.08 ng mL⁻¹ [55].
- Label-Free Optical Aptasensors: A smartphone-based fiber-optic aptasensor was created for the label-free detection of Plasmodium falciparum glutamate dehydrogenase (PfGDH), a malaria biomarker. The system functionalized a gold-sputtered optic fiber with a specific aptamer and detected binding events through changes in light signal, achieving a limit of detection of 264 pM using only 175 μL of sample [56].
Table 2. Comparison of smartphone-based digital assay platforms.
| Assay Platform | Detection Principle | Analyte | Key Performance Metric |
|---|---|---|---|
| Fluorescence Microscope [53] | Single-molecule localization (DNA-PAINT) | Ebola RNA | Single-molecule sensitivity, 84 nm resolution |
| Electrochemical Immunosensor [55] | Label-free electrochemical impedance | Carcinoembryonic Antigen (CEA) | LOD: 0.08 ng mL⁻¹, Linear range: 0.1-5.0 ng mL⁻¹ |
| Fiber-Optic Aptasensor [56] | Label-free reflectance | P. falciparum Glutamate Dehydrogenase | LOD: 264 pM, Sample volume: 175 μL |
Experimental Protocols: From Single Molecules to Super-Resolution
Sample Preparation for DNA Origami Imaging
The following protocol is adapted from the methodology used to validate the smartphone microscope [53] [57].
- Substrate Cleaning: Cut quartz coverslips to size (e.g., 49 × 16.3 mm) using a laser cutter. Clean by soaking in acetone for 1 hour, followed by rinsing with isopropanol and a second 1-hour soak in fresh isopropanol. Rinse thoroughly with Milli-Q water and dry with a stream of compressed nitrogen. Finally, bake on a hot plate at 100°C for 5 minutes to remove residual water [57].
- Grid Fabrication (for correlation): To facilitate correlation with high-end microscopes, create fiducial markers on the substrate. Using photolithography, pattern a grid onto the quartz. Deposit a 12 nm-thick chromium layer using a sputtering system. Perform a lift-off process to remove excess metal, leaving a defined grid [57].
- Surface Passivation and Functionalization: Treat the substrate with oxygen plasma to clean and activate the surface. Incubate with a passivation buffer (e.g., containing biotin-PEG) to prevent non-specific binding of biomolecules.
- Sample Immobilization: Incubate the functionalized substrate with the sample of interest. For DNA origami structures, this involves using constructs with biotin modifications that bind to a streptavidin-coated surface. Dilute the sample to a suitable density for single-molecule microscopy [53].
Workflow for Smartphone Super-Resolution Imaging
The experimental workflow for conducting super-resolution imaging with the smartphone microscope is outlined below and in Figure 2.
Figure 2. Workflow for smartphone-based super-resolution imaging (SMLM). The process involves sample preparation, optical setup, data acquisition of blinking fluorophores, and computational image reconstruction.
- Microscope Setup: Assemble the modular stages. Secure the smartphone in the silicone supports. Place the sample into the holder on the sample stage [53].
- Focusing and Alignment: Use the integrated white LED for initial coarse focusing and sample positioning. Turn off the LED and unblock the laser path. Fine-tune the alignment screws on the laser and objective stages to achieve TIR or HILO illumination, which is observed as a thin, bright line at the sample interface [53].
- Data Acquisition: Using a custom application, record a time-lapse video of the fluorescence emission. For DNA-PAINT, this involves acquiring thousands of frames (e.g., with 100 ms exposure time) to capture the stochastic "blinking" of binding events [53].
- Computational Analysis (Single-Molecule Localization):
- Pre-processing: Filter frames to identify those containing well-isolated, single-molecule emission spots.
- Localization: For each spot in each frame, computationally determine the precise center with nanometer precision using algorithms like maximum likelihood estimation or Gaussian fitting.
- Drift Correction and Merging: Compensate for stage drift and merge all localized positions from all frames into a single, high-resolution image [53] [52].
The Scientist's Toolkit: Essential Research Reagents and Materials
Successful implementation of these advanced assays requires a suite of specialized reagents and materials. The following table details key components used in the featured research.
Table 3. Essential research reagents and materials for smartphone-based super-resolution and digital assays.
| Item Name | Function/Description | Application Example |
|---|---|---|
| DNA Origami Structures | Programmable, nanoscale scaffolds for precise positioning of fluorophores and biomolecules. | Model system for validating single-molecule detection and resolution [53]. |
| ATTO 542 & ATTO 647N | Bright, photostable fluorescent dyes with well-characterized excitation/emission profiles. | Single-molecule fluorophores for detection and DNA-PAINT experiments [53]. |
| Biotin-PEG Passivation Buffer | A self-assembled monolayer that minimizes non-specific binding of proteins and nucleic acids to surfaces. | Essential for preparing clean samples for single-molecule imaging [53]. |
| Chromium Sputtering Target | Source for depositing thin, conductive metal films onto substrates. | Creating fiducial marker grids on quartz substrates for image correlation [57]. |
| Screen-Printed Carbon Electrode (SPCE) | Low-cost, disposable electrochemical cell substrate. | Base transducer for smartphone-based electrochemical immunosensors [55]. |
| Graphene Oxide (GO) / Carbon Nanotubes (CNTs) | Nanostructured carbon materials that provide high surface area and excellent electrical conductivity. | Used in layer-by-layer assemblies to amplify electrochemical signal [55]. |
| U-Bent Plastic Optic Fiber | Waveguide that enhances interaction between light and the surface coating. | Probe for label-free aptasensing; bending increases sensitivity to surface binding events [56]. |
The integration of smartphone technology with super-resolution microscopy and digital bioassays marks a significant leap toward democratizing high-precision analytical capabilities. These portable, low-cost platforms deliver performance that rivals or even surpasses that of traditional, expensive laboratory instruments in specific applications, adhering to the principles of GAC [50]. For researchers and drug development professionals, this convergence opens new avenues for point-of-care diagnostics, field-deployable analytics, and high-throughput screening with single-molecule sensitivity [53] [54] [51]. Future developments will likely be driven by the convergence of smartphones with smarter assays, advanced microfluidics, and apps powered by machine learning and artificial intelligence, promising a future for molecular analysis that is both powerful and universally accessible [51].
Overcoming Implementation Challenges in Real-World Settings
The integration of optical detection methods with smartphone-based Lab-on-a-Chip (LoC) systems represents a paradigm shift in portable molecular analysis, offering the potential to democratize analytical capabilities globally [19]. Smartphones provide a uniquely integrated package of high-resolution cameras, powerful processors, and connectivity features that can transform them into sophisticated analytical instruments [19]. However, the translation of these systems from research prototypes to reliable field-deployable tools faces three critical bottlenecks: precise sensor calibration, compensation for environmental variability, and robust signal processing of complex optical data. This technical guide examines these challenges within the context of optical detection principles and provides structured methodologies to overcome them, enabling researchers to develop more accurate, reliable, and reproducible smartphone-based LoC systems.
Bottleneck 1: Sensor Calibration and Quantitative Accuracy
Sensor calibration establishes the fundamental relationship between raw sensor outputs and meaningful quantitative measurements, serving as the foundation for all subsequent analysis in smartphone-based optical detection.
Core Calibration Principles and Methodologies
The primary goal of sensor calibration is to convert device-dependent signals (e.g., pixel intensity, voltage readings) into analyte concentrations or specific physicochemical properties. In optical sensing, this typically involves measuring the system's response to known standards to build a calibration curve. For absorbance-based measurements, this follows the Beer-Lambert law, where absorbance (A) is calculated as A = -log(It/I0), with It representing transmitted light intensity and I0 incident light intensity [58]. The precise determination of I0 is critical and requires measurement of a blank reference under identical conditions.
For fluorescence-based detection, calibration involves correlating emission intensity with analyte concentration, often requiring additional corrections for excitation source fluctuations and potential inner-filter effects at higher concentrations. Smartphone cameras typically capture signals as RGB (Red, Green, Blue) values, which must be meticulously mapped to quantitative measurements through appropriate color space transformations and channel selection optimized for the specific assay chemistry [19].
Experimental Protocol: Calibration Curve Development
Materials Required:
- Smartphone with camera and dedicated analysis app
- LoC device with integrated optical components
- Standard solutions of known analyte concentrations
- Cuvettes or microfluidic chambers compatible with the LoC device
- Stable light source (if not integrated)
Procedure:
- Prepare a series of standard solutions covering the expected dynamic range of the assay.
- For each standard, including a blank, load into the measurement chamber and acquire images using fixed smartphone camera settings (ISO, exposure, white balance).
- Extract pixel intensity values from defined regions of interest using image analysis software.
- Convert raw intensities to analytical signals (e.g., absorbance, fluorescence ratio).
- Plot analytical signal versus concentration and perform regression analysis.
- Validate the calibration model with independent standard samples.
Table 1: Quantitative Performance Metrics for Optical Sensor Calibration
| Sensor Type | Linear Dynamic Range | Limit of Detection (LOD) | Limit of Quantification (LOQ) | Reference |
|---|---|---|---|---|
| Ga³⁺ Optical Sensor | 6.25 × 10⁻⁹ to 3.75 × 10⁻⁶ M | 1.75 × 10⁻⁹ M | 6.00 × 10⁻⁹ M | [59] |
| Microplastic Optical Sensor | N/A | Visually identified particles | N/A | [58] |
| Smartphone-based Electrochemical | Pico- to femtomolar | Pico- to femtomolar | N/A | [60] |
Figure 1: Comprehensive sensor calibration workflow for smartphone-based optical detection systems
Advanced Calibration Techniques
For complex multi-analyte detection, multivariate calibration approaches such as Principal Component Regression (PCR) or Partial Least Squares (PLS) can effectively handle spectral overlaps and matrix effects. Recent advances incorporate machine learning algorithms that can learn non-linear relationships between complex optical signatures and analyte concentrations, particularly useful for direct analysis of heterogeneous samples [58]. Additionally, internal standard-based calibration methods, where a reference signal is incorporated directly into the assay, can compensate for instrumental drift and environmental fluctuations in field deployments.
Bottleneck 2: Environmental Variability and Interference
Environmental factors represent a significant challenge for reliable smartphone-based optical detection outside controlled laboratory settings, requiring systematic characterization and compensation strategies.
Ambient Light Conditions: Uncontrolled lighting represents the most significant variable in field deployments, causing fluctuating background signals and reduced signal-to-noise ratios [58]. This includes variations in intensity, spectral composition, and directionality of ambient light.
Temperature Effects: Temperature fluctuations impact reaction kinetics, optical properties of materials, and smartphone camera performance, potentially leading to signal drift.
Sample Matrix Effects: Complex real-world samples may contain interferents that cause scattering, absorption at overlapping wavelengths, or non-specific binding, particularly in biological and environmental samples [61].
Physical Variations: Inconsistent sample volume, positioning, and meniscus effects in microfluidic channels introduce measurement variability.
Experimental Protocol: Environmental Robustness Testing
Objective: Systematically evaluate and mitigate the impact of environmental variables on assay performance.
Materials:
- Environmental chamber (for controlled temperature/humidity)
- Light sources with varying spectral characteristics
- Samples with known interferents
- Portable housing/shielding prototypes
Procedure:
- Temperature Testing: Measure assay response across expected operational temperature range (e.g., 15-35°C) using constant analyte concentration.
- Lighting Condition Testing: Acquire measurements under various lighting conditions (direct sunlight, shade, artificial light) with and without shielding.
- Interference Testing: Spike samples with potential interferents at biologically/environmentally relevant concentrations.
- Sample Volume Testing: Evaluate signal variation with deliberate changes in sample volume or positioning.
- Data Analysis: Quantify the impact of each variable on measurement accuracy and precision.
Table 2: Environmental Interference and Mitigation Strategies in Optical Detection
| Interference Type | Impact on Signal | Compensation Strategies | Effectiveness |
|---|---|---|---|
| Ambient Light Fluctuation | Increased background, reduced contrast | Physical shielding, optical filters, reference normalization [58] | High with proper implementation |
| Temperature Variation | Signal drift, altered kinetics | Temperature stabilization, calibration curves at multiple temperatures | Moderate to high |
| Sample Matrix Effects | Non-specific signal, quenching | Sample purification, background subtraction, selective recognition elements [59] | Variable depending on complexity |
| Physical Positioning | Signal intensity variation | Alignment features, flow control, internal standards | High with engineered solutions |
Engineering Solutions for Environmental Compensation
Effective mitigation requires both hardware and computational strategies. Physical shielding and optical filters significantly reduce ambient light interference, as demonstrated in microplastic detection systems that use enclosed measurement chambers [58]. For temperature sensitivity, incorporating temperature sensors allows for mathematical compensation or active temperature control in more advanced systems.
From a computational perspective, background subtraction methods using reference regions or dual-wavelength measurements can effectively compensate for many environmental variables. The development of environmental interference models enables predictive compensation, where the system automatically adjusts measurements based on detected environmental conditions.
Figure 2: Environmental interference sources and mitigation pathways in smartphone-based optical detection
Bottleneck 3: Signal Processing and Data Analysis
Advanced signal processing transforms raw optical data into reliable, actionable information, addressing the inherent noise and complexity of smartphone-based detection systems.
Signal Processing Framework for Optical Data
The signal processing pipeline for smartphone-based optical detection typically involves multiple stages:
- Pre-processing: Noise reduction, flat-field correction, and image registration
- Feature Extraction: Identification and quantification of relevant optical signatures
- Data Transformation: Conversion to analytical domains (e.g., absorbance, fluorescence intensity)
- Multivariate Analysis: Handling of complex, multi-wavelength data
- Classification/Quantification: Final interpretation using calibrated models
For smartphone cameras, which typically capture RGB color space images, sophisticated processing is often required to extract quantitative information. This may involve conversion to other color spaces (e.g., HSV, Lab) that better separate chromaticity from intensity or the development of customized spectral unmixing algorithms to resolve overlapping signals [19].
Experimental Protocol: Multimodal Data Integration
Objective: Implement and validate a processing pipeline for multimodal optical data from smartphone-based detection systems.
Materials:
- Smartphone with customized analysis app
- Dataset of multi-channel optical measurements
- Computing environment for algorithm development (e.g., Python, MATLAB)
- Validation samples with known properties
Procedure:
- Data Acquisition: Collect images under standardized conditions across multiple color channels or with different optical filters.
- Pre-processing: Apply flat-field correction using reference images, implement noise reduction filters (e.g., Gaussian blur, median filtering), and perform image alignment if multiple measurements are combined.
- Region of Interest (ROI) Selection: Automatically identify relevant analysis regions while excluding artifacts or air bubbles.
- Feature Extraction: Calculate relevant features such as mean intensity, texture parameters, or spectral ratios from each ROI.
- Data Fusion: Combine features from multiple channels or modalities using appropriate weighting or multivariate methods.
- Model Development: Train classification or regression models using extracted features and known sample properties.
- Validation: Evaluate model performance using cross-validation or independent test sets.
Advanced Computational Approaches
Machine learning and artificial intelligence represent powerful tools for addressing complex signal processing challenges in smartphone-based detection. Supervised learning approaches, such as Support Vector Machines (SVM), have demonstrated high accuracy in classifying functional cell states based on multimodal optical signatures [62]. For spectral analysis, principal component analysis (PCA) and other dimensionality reduction techniques can identify latent variables that correlate with analyte concentrations while reducing noise.
Deep learning approaches, particularly convolutional neural networks (CNNs), offer significant potential for direct image-based analysis without manual feature engineering, learning optimal representations directly from raw pixel data. These approaches are particularly valuable for complex detection tasks such as identifying microplastics in environmental samples or classifying cell states in biomedical applications [58].
Table 3: Signal Processing Methods for Smartphone-Based Optical Detection
| Processing Method | Application Context | Key Advantages | Implementation Considerations |
|---|---|---|---|
| Support Vector Machines (SVM) | Classification of cell death modes [62] | Effective in high-dimensional spaces, memory efficient | Requires careful kernel selection and parameter tuning |
| Principal Component Analysis (PCA) | Spectral unmixing, dimensionality reduction | Reduces noise, identifies latent variables | Linear method, may miss complex nonlinear relationships |
| Convolutional Neural Networks (CNN) | Image-based classification, feature extraction | Automatic feature learning, state-of-the-art performance | Requires large labeled datasets, computationally intensive |
| Multimodal Data Fusion | Combining multiple contrast mechanisms [62] | Leverages complementary information, improved accuracy | Registration challenges, heterogeneous data types |
The Scientist's Toolkit: Essential Research Reagents and Materials
Successful development of smartphone-based optical detection systems requires careful selection of recognition elements, materials, and instrumentation components.
Table 4: Essential Research Reagents and Materials for Smartphone-Based Optical LoC Development
| Category | Specific Examples | Function in Optical Detection | Implementation Considerations |
|---|---|---|---|
| Biological Recognition Elements | Enzymes, antibodies, aptamers, nucleic acids [60] | Provide molecular specificity through selective binding | Stability, immobilization method, non-specific binding |
| Optical Materials | Chromoionophores (e.g., ETH-5294) [59], fluorescent dyes, quantum dots | Transduce molecular recognition into optical signals | Photostability, compatibility with excitation/emission filters |
| Nanomaterials | Gold nanoparticles, graphene oxide, carbon dots [60] | Enhance signal intensity, improve detection limits | Biocompatibility, functionalization chemistry, aggregation |
| Substrate Materials | Polyvinyl chloride (PVC) membranes [59], PDMS, glass | Provide support for recognition elements, optical clarity | Autofluorescence, chemical compatibility, surface chemistry |
| Light Sources | LEDs, laser diodes | Provide excitation for fluorescence or illumination for absorbance | Spectral characteristics, intensity stability, power requirements |
| Optical Components | Filters, lenses, optical fibers [61] | Control light path, select specific wavelengths | Alignment requirements, transmission efficiency, stray light |
Addressing the critical bottlenecks of sensor calibration, environmental variability, and signal processing is essential for advancing smartphone-based optical LoC systems from research prototypes to reliable analytical tools. Through systematic calibration protocols, engineered solutions for environmental compensation, and sophisticated signal processing approaches, researchers can significantly enhance the reliability and performance of these portable detection platforms.
Future advancements will likely incorporate increasingly intelligent systems that leverage machine learning not only for data analysis but also for real-time optimization of acquisition parameters and automatic quality control. The integration of multimodal detection approaches, combining optical with electrochemical methods [60], offers complementary information that can improve accuracy in complex samples. Additionally, the development of standardized validation frameworks and reference materials will be crucial for establishing credibility and facilitating adoption across healthcare, environmental monitoring, and food safety applications.
As these technologies mature, smartphone-based optical detection systems have the potential to fundamentally transform analytical capabilities, making sophisticated molecular analysis accessible outside traditional laboratory settings and supporting distributed monitoring networks for global health and environmental protection.
In smartphone-based Lab-on-Chip (LoC) research, the principles of optical detection are paramount. These portable, affordable systems promise to revolutionize point-of-care diagnostics and biomedical research, yet their performance is highly dependent on the quality of data acquisition. Unlike controlled laboratory environments, smartphone-based systems operate under variable and suboptimal conditions, making robust strategies for lighting control and image analysis algorithms essential. This technical guide explores these core challenges within the context of a broader thesis on optical detection methods, providing researchers and drug development professionals with methodologies to enhance the reliability and accuracy of their portable diagnostic systems.
The Critical Role of Controlled Illumination
Inconsistent lighting is a primary source of error in optical detection, introducing artifacts, reducing contrast, and compromising quantitative analysis. Controlling this variable is a foundational step toward robust data acquisition.
The Flash-No-Flash (FNF) Protocol
The FNF acquisition protocol is a powerful method to stabilize the impact of ambient lighting. This approach involves the near-simultaneous capture of two images: one with a strong, integrated artificial light source ("Flash") and one with ambient light only ("No-Flash"). The difference between these images represents the scene as if illuminated only by the controlled artificial source, effectively canceling out the variable ambient light [63].
Experimental Protocol for FNF Implementation:
- Hardware Setup: Integrate a white light LED array controllable via the smartphone's software (e.g., through a Bluetooth-enabled microcontroller). The camera's exposure time should be set to the lowest possible value (e.g., 20 μs) to minimize ambient light contribution [63].
- Image Pair Acquisition: Program the system to alternate the LED trigger between consecutive frames. The system must automatically identify valid FNF pairs by comparing the average brightness of successive images against a pre-defined threshold [63].
- Image Processing: Subtract the "No-Flash" image from the "Flash" image on a per-pixel basis. Correct for artifacts by excluding overexposed pixels in the "Flash" image from subtraction and setting any resulting negative pixel values to zero to produce a valid RGB image for analysis [63].
Flickerless LED Illumination
Pulse-Width Modulation (PWM), a common method for dimming LEDs, can create significant image artifacts. With a rolling shutter CMOS sensor (common in smartphones), the changing light during the sensor's scan results in banding across the image. Even with a global shutter, unsynchronized PWM can cause frame-to-frame brightness variability [64].
Flickerless Dimming Methodology:
- Implementation: Replace PWM-driven LEDs with a DC/DC buck or boost LED driver in tracking mode. The LED current is controlled by an adjustable voltage reference from an I2C-controlled Digital-to-Analog Converter (DAC), providing linear brightness control.
- Synchronization: To achieve stable illumination, the LED driver should be tightly coupled with the image acquisition process. This can be done by:
- Using the sensor's frame synchronization signal to gate the LED driver.
- Implementing accurate timing synchronization between the light parameters and image acquisition trigger using programmable logic [64].
Table 1: Comparison of Illumination Control Strategies
| Strategy | Core Principle | Key Advantage | Implementation Complexity | Best Suited For |
|---|---|---|---|---|
| Flash-No-Flash (FNF) [63] | Computational subtraction of ambient light | Effectively removes all ambient light effects | High (requires hardware control & image processing) | Field applications with highly variable ambient light |
| Flickerless LED [64] | Analog current control & host synchronization | Eliminates temporal artifacts from PWM dimming | Medium (requires specific driver hardware) | All applications requiring stable, adjustable illumination |
| Intelligent Luminance Control [65] | Closed-loop feedback using a wearable camera | Automatically maintains optimal lighting conditions | High (requires feedback system & LEEM benchmarks) | Desk-based, long-term monitoring applications |
Optimizing Image Analysis Algorithms
With controlled illumination producing consistent input data, the next step is to deploy image analysis algorithms that are both robust and efficient enough to run on smartphone-level hardware.
Adaptive, Morphology-Independent Segmentation
Deep learning offers high accuracy but demands significant computational resources and large, annotated datasets. For many LoC applications, adaptive algorithms that do not require extensive training are more practical.
Algorithmic Workflow: The Quantella platform demonstrates an effective adaptive pipeline for cell analysis [66]:
- Image Enhancement: An initial step improves raw data quality, enhancing boundaries and contrast.
- Multi-Exposure Fusion & Thresholding: The algorithm uses multi-weight-map analysis and thresholding to accurately segment cells.
- Morphological Filtering: This step refines the segmentation, distinguishing individual cells even in densely clustered samples without being dependent on cell-specific morphology or user-defined parameters [66].
This approach has been validated across diverse cell types (suspension, adherent, and primary cells like RBCs), achieving over 90% accuracy in cell identification and deviations of less than 5% compared to flow cytometry, while analyzing over 10,000 cells per test [66].
Smartphone-Based TLC Analysis
The analysis of Thin-Layer Chromatography (TLC) plates is a common application in pharmaceutical quality screening. A smartphone-based method can make this quantitative analysis portable and accessible.
Experimental Protocol for TLC Analysis [67]:
- Imaging Setup: A custom-made UV imaging box (e.g., 25 cm x 15 cm x 15 cm cardboard) is used to standardize imaging. A rear-facing smartphone camera captures images through a hole in the lid, with the TLC plate inserted via a front slit.
- Image Processing Algorithm:
- Channel Extraction & Inversion: The captured RGB image is cropped to the region of interest, and the green channel is extracted, inverted, and normalized.
- Noise Reduction & Thresholding: A 2D Gaussian filter (5x5 kernel) smooths the image. Subsequent dilation and binary thresholding create a binary matrix.
- Contour Detection: OpenCV's contour detection function identifies the spots, and the moment function locates their centers for calculating retention factors (Rf) and quantifying spot intensity [67].
This method demonstrated high consistency with ImageJ software and successfully analyzed metformin samples from local pharmacies, identifying 15 of 16 samples as containing acceptable drug levels [67].
Integrated Workflow for Robust Optical Detection
The described strategies for lighting control and algorithm optimization form a cohesive workflow for enhancing data acquisition in smartphone-based LoC systems. The diagram below illustrates the logical relationships and sequential stages of this integrated approach.
The Researcher's Toolkit: Essential Materials and Reagents
Implementing the described strategies requires a set of key hardware and software components. The following table details essential research reagent solutions and their functions in establishing a robust smartphone-based optical detection system.
Table 2: Key Research Reagent Solutions for Smartphone-Based LoC Systems
| Item Name | Function / Role | Implementation Example |
|---|---|---|
| Smartphone with Programmable Camera | The primary image sensor and computational unit. | Oppo Reno 10× Zoom used in Quantella platform for cell analysis [66]. |
| Controllable LED Array | Provides a stable, artificial light source for illumination control. | Integrated white LED array for FNF imaging; UV LED for TLC plate excitation [63] [67]. |
| Microcontroller (e.g., Arduino) | Interfaces between smartphone and hardware, enabling control of pumps and lights. | Arduino used to control a piezoelectric pump's flow rate via Bluetooth in Quantella [66]. |
| Open-Source Computer Vision Library (OpenCV) | Provides core algorithms for image processing and analysis. | OpenCV V3.42 used in the TLC Analyzer app for contour detection and analysis [67]. |
| Optofluidic Flow Cell | Presents a consistent, known volume for sample imaging. | A single-channel flow cell (100 μm wide, similar to a hemocytometer) used in Quantella [66]. |
| Pre-coated TLC Plates | The stationary phase for separation in chromatographic drug analysis. | Silica gel 60 F254 plates used for metformin separation [67]. |
| Standard Color Card | Provides a color benchmark for calibration and color deviation analysis. | Pasted on the margin of a display to aid intelligent luminance control systems [65]. |
Achieving robust data acquisition in smartphone-based LoC research is a multifaceted challenge addressed by tackling both the physical acquisition environment and the computational analysis pipeline. As demonstrated, controlling illumination through protocols like FNF and flickerless lighting directly mitigates the primary source of noise—variable ambient light. Subsequently, employing optimized, adaptive image analysis algorithms ensures that accurate, quantitative data can be extracted efficiently on modest hardware. Together, these strategies form a cohesive methodology that enhances the reliability and scalability of optical detection methods, pushing the frontier of accessible and precise point-of-care diagnostics and biomedical research.
Enhancing Sensitivity and Reproducibility with Nanomaterials and Advanced Signal Processing
The integration of optical detection methods into smartphone-based lab-on-a-chip (LoC) systems represents a paradigm shift in point-of-care diagnostics, environmental monitoring, and food safety testing. These portable platforms leverage the ubiquitous nature of smartphones, which provide substantial computational power, high-resolution cameras, and connectivity features in a globally accessible format [19]. A critical challenge, however, lies in achieving the sensitivity and reproducibility typically associated with bulky, centralized laboratory equipment. This is where the synergistic combination of engineered nanomaterials and advanced digital signal processing becomes transformative. Nanomaterials directly enhance the physical and optical interactions at the sensor interface, amplifying signals and improving stability. Concurrently, machine learning (ML) and artificial intelligence (AI) algorithms process the complex, nanomaterial-enhanced signals to extract robust, quantitative data, thereby overcoming variability and noise. This technical guide examines the principles and methodologies underpinning this convergence, providing a framework for developing next-generation smartphone-based optical biosensors characterized by high sensitivity and exceptional reproducibility.
Nanomaterial Engineering for Enhanced Optical Sensitivity
The strategic incorporation of nanomaterials into the sensing interface is foundational to enhancing signal intensity and stability. Their unique physicochemical properties, derived from high surface-to-volume ratios and quantum effects, directly address the sensitivity limitations of conventional assays.
Key Nanomaterials and Their Signal Amplification Mechanisms
Table 1: Functional Nanomaterials for Enhancing Optical Biosensor Sensitivity
| Nanomaterial | Core Properties | Optical Mechanism | Impact on Sensitivity |
|---|---|---|---|
| Gold Nanoparticles (AuNPs) | Biocompatibility, tunable LSPR, strong scattering [68] [69] | LSPR, MEF, catalytic activity [69] | Enhances fluorescence quantum yield, enables label-free detection via refractive index shifts [69]. |
| Silver Nanoparticles (AgNPs) | Superior conductivity, high plasmon resonance frequency [69] | Intense LSPR fields, MEF [69] | Provides greater fluorescence enhancement than AuNPs; enables single-molecule detection [69]. |
| Graphene Oxide (GO) | Large 2D surface area, oxygen functional groups (-OH, -COOH) [60] | Fluorescence quenching (FRET), adsorption [69] | Lowers background noise via efficient FRET quenching; enables signal-on detection [69]. |
| Quantum Dots (QDs) | Size-tunable emission, high photostability, bright fluorescence [68] | Narrow, symmetric photoluminescence [68] | Provides stable, multiplexed fluorescence signals resistant to photobleaching [68]. |
| MXenes | Tunable surface chemistry, strong plasmonic/catalytic activity [68] [69] | Enhanced adsorption, catalytic signal amplification [69] | Improves electrode conductivity and biomolecule immobilization in electrochemical-optical systems [68]. |
Experimental Protocol: Surface Functionalization for Reproducibility
A critical step in leveraging nanomaterials is their consistent and stable functionalization with biorecognition elements. The following protocol for functionalizing gold nanoparticles (AuNPs) with thiolated DNA aptamers ensures high binding efficiency and low non-specific adsorption [70] [69].
- Nanoparticle Preparation: Synthesize or acquire spherical AuNPs (e.g., 20-40 nm diameter) characterized by UV-Vis spectroscopy to confirm the LSPR peak position and particle size uniformity [69].
- Surface Cleaning: Incubate the AuNP solution with a 0.1% w/v solution of Triton X-100 for 15 minutes, followed by centrifugation (14,000 rpm, 20 minutes) and resuspension in deionized water. This removes residual surfactants and contaminants.
- Probe Immobilization: a. Prepare a 1 µM solution of thiolated DNA aptamer in a 10 mM Tris-HCl buffer (pH 8.0) containing 1 mM EDTA. b. Add 10 µL of 100 mM Tris(2-carboxyethyl)phosphine (TCEP) to 100 µL of the aptamer solution and incubate for 1 hour at room temperature to reduce disulfide bonds. c. Mix the reduced aptamer solution with the purified AuNP solution at a ratio of 200 DNA strands per nanoparticle. d. Allow the conjugation to proceed for 16-24 hours at room temperature with gentle shaking.
- Backfilling and Stabilization: a. Add a 1000-fold molar excess of 6-mercapto-1-hexanol (MCH) relative to the aptamer concentration and incubate for 4 hours. This step passivates the uncovered gold surface, minimizes non-specific binding, and orientates the aptamers upright. b. Centrifuge the functionalized AuNPs (14,000 rpm, 20 minutes) and carefully remove the supernatant. c. Resuspend the pellet in a suitable storage buffer (e.g., 10 mM PBS with 0.1% BSA, pH 7.4).
- Validation: Characterize the functionalized AuNPs using dynamic light scattering (DLS) to confirm an increase in hydrodynamic diameter and UV-Vis spectroscopy to ensure no aggregation has occurred (maintained LSPR peak).
Smartphone-Based Optical Detection Modalities
Smartphones serve as the central hub for optical signal capture and initial processing. Their CMOS cameras are the primary detectors, adapted for various optical readout modalities.
Diagram 1: Core smartphone optical detection workflow.
Critical Optical Configurations and Methodologies
Colorimetric Detection: This method relies on target-induced color changes, often amplified by the intense localized surface plasmon resonance (LSPR) of noble metal nanoparticles. The smartphone camera captures an image of the sensor area under consistent illumination. The RGB (Red, Green, Blue) values are extracted using an onboard app and correlated to analyte concentration via a pre-calibrated curve. Aggregation of AuNPs, for instance, causes a visible color shift from red to blue, providing a robust visual readout [19] [69].
Fluorescence Detection: Fluorophores, such as quantum dots or organic dyes, are excited by an external light source (e.g., a low-cost LED). The smartphone camera, often fitted with an emission filter to block the excitation light, captures the emitted fluorescence. Nanomaterials like AgNPs or Au nanostars can be used to create metal-enhanced fluorescence (MEF), boosting the signal intensity by up to 1500-fold, which dramatically lowers the limit of detection [69]. The smartphone app analyzes the intensity or color of the fluorescence.
Surface-Enhanced Raman Scattering (SERS): This technique provides molecular "fingerprinting" through inelastic light scattering, which is dramatically amplified by plasmonic nanostructures. While traditionally requiring complex spectrometers, smartphone-based SERS systems are emerging. They use a simplified optical setup where the smartphone camera, coupled with a laser diode and a notch filter, captures the unique Raman spectrum, which is then decoded by ML algorithms to identify and quantify the analyte [68] [71].
Advanced Signal Processing for Data Reproducibility
The raw data captured by the smartphone is often corrupted by noise from various sources, including uneven illumination, sensor noise, and environmental fluctuations. Advanced signal processing, particularly AI and ML, is essential to transform this variable data into reproducible, quantitative results.
AI and Machine Learning Integration Workflow
Diagram 2: AI-powered signal processing for reproducibility.
Key Algorithms and Implementation Protocols
Table 2: Advanced Signal Processing Techniques for Enhanced Reproducibility
| Processing Stage | Algorithm/Technique | Function | Example Implementation |
|---|---|---|---|
| Signal Denoising | Chebyshev Type-I Filter [72] | Removes high-frequency noise from inertial and spectral time-series data. | Applied to raw accelerometer/gyroscope data before activity recognition; improves signal-to-noise ratio. |
| Feature Selection | Boruta Algorithm [72] | Identifies all-relevant features from extracted data, reducing dimensionality. | Selects the most informative color channels, intensity statistics, and texture features from a smartphone-captured image. |
| Data Optimization | Particle Swarm Optimization (PSO) [72] | Iteratively refines feature vectors to find an optimal set for model training. | Optimizes the input vector for a recurrent neural network (RNN) to maximize classification accuracy for a colorimetric assay. |
| Intelligent Modeling | Recurrent Neural Networks (RNNs) [72] | Models temporal dependencies in sequential sensor data. | Used for tracking dynamic signal changes in real-time monitoring, adapting to evolving signal patterns. |
| Intelligent Modeling | Convolutional Neural Networks (CNNs) [71] | Extracts spatial features from image-based data (e.g., spots, color gradients). | Automatically identifies and quantifies fluorescence signals from a microarray image, ignoring illumination artifacts. |
| Image Analysis | AI-Assisted Image Reconstruction [68] [71] | Reconstructs super-resolution images or corrects for optical aberrations. | Enables resolution beyond the diffraction limit of the smartphone camera optics when paired with nanoscale light manipulation. |
Protocol: Implementing a Chebyshev Filter for Signal Denoising
This protocol is crucial for preprocessing data from smartphone inertial sensors or for smoothing temporal optical signals [72].
- Signal Acquisition: Collect the raw signal from the smartphone sensor (e.g., accelerometer, or a photodiode measuring light intensity over time) at a specified sampling rate (e.g., 100 Hz).
- Parameter Definition: Define the filter parameters. For a low-pass Chebyshev Type-I filter:
- Passband frequency (Fp): The highest frequency to pass unchanged (e.g., 5 Hz for slow-varying signals).
- Stopband frequency (Fs): The frequency where attenuation begins (e.g., 10 Hz).
- Passband ripple (Rp): The maximum allowed ripple in the passband (e.g., 0.1 dB).
- Stopband attenuation (Rs): The minimum attenuation required in the stopband (e.g., 40 dB).
- Filter Design: Use a computational library (e.g.,
scipy.signalin Python) to design the filter.N, Wn = signal.cheb1ord(Fp, Fs, Rp, Rs, fs=sampling_rate)b, a = signal.cheby1(N, Rp, Wn, 'low')
- Signal Filtering: Apply the filter to the raw signal using the
signal.filtfilt(b, a, raw_signal)function. Zero-phase filtering (filtfilt) is preferred as it avoids distorting the signal's phase. - Validation: Plot the original and filtered signals to visually confirm noise reduction without significant distortion of the signal's fundamental shape. Calculate metrics like the Signal-to-Noise Ratio (SNR) before and after filtering.
The Scientist's Toolkit: Essential Research Reagent Solutions
Table 3: Key Reagents and Materials for Developing Smartphone-Based Nanobiosensors
| Item | Function/Description | Application Example |
|---|---|---|
| Thiolated DNA Aptamers | Biorecognition elements that bind to specific targets (proteins, small molecules); thiol group allows for covalent binding to gold surfaces [60] [69]. | Functionalization of AuNPs for colorimetric or electrochemical-optical detection of biomarkers like CEA or viruses [69]. |
| Polyethylene Glycol (PEG) | A polymer spacer used in surface functionalization to minimize non-specific adsorption of biomolecules, improving signal-to-noise ratio and reproducibility [70] [69]. | Backfilling on sensor surfaces after probe immobilization to create a bio-inert background. |
| Tris(2-carboxyethyl)phosphine (TCEP) | A reducing agent that cleaves disulfide bonds without the need for purification, activating thiolated DNA for conjugation [69]. | Pre-treatment of thiolated aptamers or antibodies before immobilization on gold nanostructures. |
| Microfluidic Chip | A device with micron-sized channels that automates fluid handling, enabling precise control over sample and reagent volumes, which is critical for assay reproducibility [68] [60]. | Integrated with a nanobiosensor for automated sample preparation, mixing, and delivery to the detection zone in a smartphone LoC device. |
| Quantum Dots (CdSe/ZnS) | Semiconductor nanoparticles serving as photostable, multiplexable fluorescent labels with size-tunable emission wavelengths [68]. | Used as fluorescent tags in sandwich immunoassays for simultaneous detection of multiple disease biomarkers. |
| Gold Nanorods | Anisotropic gold nanoparticles with two LSPR bands (transverse and longitudinal); the longitudinal band is highly sensitive to the local environment and aggregation [69]. | Label-free LSPR biosensing; the longitudinal peak shift upon target binding is quantified using the smartphone camera. |
The integration of optical detection methods into smartphone-based Lab-on-Chip (LoC) systems represents a paradigm shift in point-of-care diagnostics. These systems leverage the ubiquitous nature of smartphones, which are equipped with highly integrated sensors, rapidly evolving computational power, and widespread user adoption [73]. For researchers and drug development professionals, the primary challenge has shifted from proving technical feasibility to bridging the critical integration gap: creating systems that seamlessly connect with existing healthcare data infrastructures and are manufacturable at scale. This whitepaper provides a technical framework for addressing these dual challenges, contextualized within the broader thesis of optical detection principles in smartphone-based LoC research. We present detailed methodologies, quantitative performance data, and implementation protocols to guide the transition from laboratory prototypes to clinically validated, commercially viable diagnostic platforms.
Core Principles of Optical Detection in Smartphone-Based LoC Systems
Smartphone-based optical detection leverages several fundamental principles of light-matter interaction. The core architecture typically involves an LoC device that interfaces with a smartphone's built-in optical sensors—primarily the camera, but increasingly including complementary sensors like ambient light sensors—to quantify analytical targets.
Technical Foundations and Modalities
The primary optical modalities employed in smartphone-based detection include:
- Colorimetric Analysis: Measures color intensity changes from biochemical reactions (e.g., enzyme-linked assays). The smartphone camera captures images which are converted from RGB to hue-saturation-value (HSV) or CIE Lab color spaces for more precise quantitative analysis [74].
- Photoluminescence Detection: Utilizes light emission from excited molecules (fluorescence, chemiluminescence). This often requires additional low-cost optical components like light-emitting diodes (LEDs) for excitation and interference filters for emission wavelength selection.
- Absorbance Spectroscopy: Quantifies the absorption of specific light wavelengths by analytes. While traditionally requiring bulky spectrometers, smartphone implementations use the camera's inherent capability to act as a rudimentary spectrophotometer.
- Light Scattering Techniques: Detects changes in light scattering properties due to particle aggregation or cell binding, useful for immunoassays and pathogen detection.
The performance of these systems is fundamentally governed by their optical design, which must balance competing demands of sensitivity, specificity, cost, and manufacturability [74]. Key technical considerations include illumination stability, optical path design, and the signal-to-noise ratio of the detection system.
Connectivity with Healthcare Systems: Technical Frameworks and Protocols
For smartphone-based LoC devices to transition from research tools to clinical assets, they must achieve seamless bidirectional data exchange with established healthcare information systems, including Electronic Health Records (EHRs), laboratory information systems (LIS), and public health reporting infrastructures.
Data Architecture and Interoperability Standards
A robust data architecture is the foundation for healthcare connectivity. The implementation requires a multi-layered approach:
- Device Layer: The smartphone application must not only perform the assay analysis but also package results with essential metadata—including device calibration status, timestamp, patient identifier (hashed), and quality control metrics.
- Integration Layer: This layer implements critical healthcare data standards. HL7 FHIR (Fast Healthcare Interoperability Resources) has emerged as the dominant standard for representing clinical data as discrete data elements (e.g.,
Observationfor a test result,Devicefor the LoC reader). DICOMweb can be utilized for storing and retrieving diagnostic images captured during the assay. - Security Layer: Data in transit must be protected using TLS 1.2 or higher. Data at rest, particularly Protected Health Information (PHI), should be encrypted using AES-256. Authentication should be implemented via OAuth 2.0, while authorization should adhere to role-based access control models.
Table 1: Quantitative Performance Metrics for Healthcare System Integration
| Integration Parameter | Target Performance Metric | Measurement Protocol |
|---|---|---|
| Data Transmission Success Rate | >99.5% | Measure end-to-end success of result delivery from app to EHR over 1,000 trials under variable network conditions. |
| Result Delivery Latency | < 60 seconds | Time interval from user confirming result upload to database write confirmation in the destination system. |
| HL7 FHIR Compliance | 100% core elements | Validation against FHIR Observation profile using official FHIR validation tools. |
| API Availability | >99.9% uptime | Monitor API endpoint using synthetic transactions from multiple geographic locations. |
Experimental Protocol: Validation of End-to-End Connectivity
Aim: To verify the reliable and secure transmission of quantitative assay results from a smartphone LoC application to a test EHR environment.
Materials:
- Smartphone with custom LoC analysis application
- Calibrated LoC device with simulated sample (e.g., dye solution)
- Test EHR sandbox with FHIR API endpoint
- Network conditioning tool (to simulate 4G/Wi-Fi)
- Data monitoring software (e.g., Wireshark, Postman)
Methodology:
- Setup: Configure the LoC application with the correct API endpoints and authentication credentials for the test EHR sandbox.
- Data Generation: Run five replicate analyses of the calibrated sample using the smartphone-LoC system. The app should generate a result in a standard unit (e.g., ng/mL) with a timestamp and a unique measurement ID.
- Transmission: Initiate result upload from the application. Use the network conditioning tool to introduce packet loss (up to 2%) and latency (up to 200ms) to simulate real-world conditions.
- Verification: Using the data monitoring software, confirm that the outbound HTTP POST request contains a valid FHIR
Observationresource in the request body. The resource must include:status:finalcode: A LOINC code representing the assay (e.g.,12541-0for a generic protein assay)valueQuantity: The numerical result, unit, and systemdevice: A reference to the device resource for the LoC reader
- Validation: Query the EHR sandbox API within 60 seconds to confirm the received result matches the transmitted result exactly. Check the audit log in the EHR to confirm the data source.
Acceptance Criterion: 100% of the five replicate results must be accurately recorded in the EHR sandbox within 60 seconds without manual intervention.
Scalable Manufacturing of Integrated Optical LoC Systems
The manufacturing leap from lab prototypes to high-volume, consistent, and reliable products demands meticulous attention to optomechanical design, assembly processes, and supply chain management.
Design for Manufacturability (DFM) in Optical Systems
Successful scaling requires converting precise optical requirements into a manufacturable optomechanical design [74]. Key strategies include:
- Tolerance Analysis and Error Budgeting: Perform simulations to allocate error budgets across all optical (lens alignment, filter tilt) and mechanical (housing warpage, adhesive shrinkage) components. This ensures final assembled performance meets specifications despite inherent manufacturing variations.
- Design Simplification: Minimize the number of optical components and unique parts. Use commercially available, off-the-shelf (COTS) lenses and filters where possible to avoid costly custom optics.
- Design for Assembly (DFA): Create designs that are easy to align and assemble. This includes incorporating self-locating features, kinematic mounts, and snap-fits to reduce the need for complex fixturing and manual adjustment.
Table 2: Key Research Reagent Solutions and Materials for Scalable LoC Manufacturing
| Material / Component | Function | Scalability Consideration |
|---|---|---|
| Injection Molded Cyclic Olefin Copolymer (COC) | Microfluidic cartridge body; excellent optical clarity, low autofluorescence. | High upfront tooling cost, but extremely low cost per part at volume (>10k units). |
| Reagent Lyophilization Beads | Stable, dry storage of assay reagents within the cartridge. | Enables long shelf-life at ambient temperatures; compatible with automated pick-and-place equipment. |
| Pressure-Sensitive Adhesive (PSA) Laminate | Seals microfluidic channels; can incorporate embedded filters or membranes. | Die-cut PSAs allow for rapid, high-throughput sealing vs. slower liquid adhesives or thermal bonding. |
| Integrated Waveguide Structures | Built-in optical paths for illumination and signal collection. | Can be monolithically fabricated into the cartridge, reducing external optical components and alignment steps. |
| Active Alignment Fixtures | Manufacturing tools that provide real-time feedback during critical assembly steps. | Crucial for achieving high production yield in systems requiring precise optical alignment [74]. |
Experimental Protocol: Validating Manufacturing Consistency
Aim: To assess the inter-device and intra-device variability of a smartphone-based LoC reader across a pilot production run of 100 units.
Materials:
- 100 LoC readers from the pilot production line
- 500 pre-qualified, lot-controlled test cartridges containing a stable fluorophore (e.g., 1 µM Fluorescein)
- Master calibration smartphone device
- Dark test chamber
Methodology:
- Setup: Place each reader and a test cartridge into the dark chamber. Use a robotic jig to ensure consistent positioning of the smartphone.
- Measurement: For each of the 100 readers, run the integrated assay protocol five times, using a fresh test cartridge each time. The protocol should automatically run and record the raw signal intensity (e.g., mean green pixel value from the camera).
- Data Analysis: Calculate the following:
- Intra-device Precision: For each reader, calculate the Coefficient of Variation (CV) across its five replicate measurements.
- Inter-device Precision: Calculate the mean signal for each reader, then calculate the CV of these 100 means.
- Accuracy/Bias: Compare the overall mean signal of the 100 readers to the mean signal obtained from the master calibration device. Express as a percentage difference.
Acceptance Criteria:
- Intra-device CV for all readers shall be < 5%.
- Inter-device CV of the mean signals shall be < 8%.
- Mean bias from the master device shall be < 10%.
Integrated Workflow and Logical Architecture
The complete integration of detection, data processing, and healthcare reporting requires a coherent logical architecture. The diagram below illustrates this end-to-end workflow.
Figure 1: End-to-End Integrated LoC System Workflow. This diagram outlines the logical flow of data and materials from sample introduction to clinical data integration, highlighting the critical handoff points between the physical device, mobile application, cloud infrastructure, and healthcare systems.
Bridging the integration gap for smartphone-based LoC devices is a multi-disciplinary challenge that extends far beyond the core optical detection science. Success hinges on the simultaneous mastery of two complex domains: the implementation of robust, standards-based healthcare data connectivity and the application of rigorous design-for-manufacturing principles to optical system engineering. The protocols and frameworks presented herein provide a concrete foundation for researchers and developers to build upon. By treating connectivity and manufacturability as first-class design requirements from the earliest stages of development, the field can accelerate the translation of these promising technologies from the research bench to the patient's hands, ultimately unlocking their full potential to revolutionize diagnostic medicine and drug development.
Performance Benchmarking and Strategic Selection of Optical Methods
The evolution of lab-on-a-chip (LOC) technologies promises to revolutionize chemical and biological analysis by making it portable, affordable, and accessible. A critical enabler of this transition is the integration of smartphones as versatile detection platforms. Their global ubiquity, integrated sensors, and powerful processing capabilities offer a shortcut to creating deployable analytical devices [19]. Within this paradigm, the choice of optical detection modality directly controls the performance and limits of detection (LOD) achievable by the system. This review provides a comparative analysis of prominent optical sensing methods—spectrophotometry, LED photometry, and camera-based imaging—within the context of smartphone-based LOC research. We synthesize recent findings to evaluate their performance metrics, detail experimental protocols for their implementation, and discuss their suitability for specific analytical scenarios, with a particular focus on the stringent demands of drug development and molecular diagnostics.
Core Optical Modalities in Smartphone-Based Detection
Optical sensing remains one of the most reliable and cost-effective methods for obtaining bio/chemical information [75]. For smartphone-based LOC systems, three primary approaches have emerged, each with distinct operational principles and hardware requirements.
1. Spectrophotometry: This laboratory-grade method measures the absorption of light by a sample across a range of wavelengths. While traditional spectrophotometers are benchtop instruments, miniaturized versions can be interfaced with smartphones for data processing and display, though often at the cost of portability [75].
2. LED Photometry (PEDD): The Paired Emitter–Detector Diode (PEDD) approach is a low-cost, highly sensitive photometric method. It typically uses a single-wavelength light-emitting diode (LED) as the source and another LED of a similar type as the light detector, operating in a charge-discharge cycle. This method is notable for its simplicity, low cost, and high performance [75].
3. Camera-Based Imaging: This method leverages the smartphone's built-in camera as a two-dimensional array detector. It can be implemented in two primary ways: Smartphone-Based Digital Image Analysis (SBDIA), which involves capturing a digital image of the sample and analyzing color-based characteristics (e.g., RGB values), and direct colorimetric analysis, where the smartphone measures light intensity emitted from or transmitted through a sample [50]. The camera can be used for a wide range of analyses, including colorimetry, fluorescence, and even microscopy [19].
Quantitative Performance Comparison
A rigorous comparative study of these three optical sensing approaches for colorimetric pH detection revealed significant differences in their performance characteristics. The following table summarizes the key sensory metrics, using spectrophotometry as the baseline for comparison [75].
Table 1: Performance comparison of optical sensing modalities for colorimetric bio/chemical detection [75].
| Performance Metric | Spectrophotometry | LED Photometry (PEDD) | Camera-Based Imaging |
|---|---|---|---|
| Measurement Range | 1x (Baseline) | 16.39x Improvement | Not Specified |
| Dynamic Range | 1x (Baseline) | 147.06x Improvement | Not Specified |
| Accuracy | 1x (Baseline) | 1.79x Improvement | Not Specified |
| Sensitivity | 1x (Baseline) | 107.53x Improvement | Not Specified |
| Limit of Detection (LOD) | Moderate | Superior (Lowest) | Lower Performance |
| Resolution | Moderate | Superior | Lower Performance |
| Key Strengths | Laboratory standard, full spectrum | High sensitivity, cost-effective, portable | Ubiquity, ease of use, rich spatial data |
| Key Limitations | Cost, size, power consumption | Single wavelength | Susceptible to ambient light, lower precision |
The data demonstrates that the LED-based PEDD system outperformed the other two methods in key sensory metrics, including sensitivity, resolution, and limit of detection, while also offering advantages in cost-effectiveness and scalability [75]. This makes it a particularly compelling solution for industrial and field applications.
Pushing the boundaries of sensitivity, recent advancements have demonstrated that smartphone-based microscopes can achieve single-molecule detection. One study developed a portable, inexpensive smartphone-based fluorescence microscope capable of direct single-molecule detection without signal amplification. This device, costing under €350, achieved a signal-to-noise ratio of 3.3 when detecting single ATTO 542 dyes on DNA origami structures and was later used for super-resolution imaging of cellular microtubule networks [53]. This represents the ultimate limit of detection and opens new possibilities for digital bioassays and point-of-care diagnostics.
Detailed Experimental Protocols
To ensure reproducibility and provide a clear technical roadmap, this section outlines the methodologies for key experiments cited in the performance comparison and for achieving state-of-the-art detection limits.
Protocol: Comparative Colorimetric Analysis
The following workflow is adapted from a systematic study comparing spectrophotometry, LED photometry, and imaging [75].
1. Sample Preparation:
- Prepare a stock solution of a colorimetric pH indicator (e.g., 50 µM Bromocresol Green in ultrapure water).
- Using controlled titration with 0.1 M HCl and 0.1 M KOH, prepare solutions spanning a pH range (e.g., pH 2–8).
- Add a consistent volume of the dye stock to each pH solution to ensure a uniform indicator concentration (e.g., 25 µM).
- Transfer aliquots (e.g., 2 mL) to standard cuvettes for analysis.
2. Reference Measurements:
- Calibrate a pH meter with standard buffers.
- Measure the pH of each solution in triplicate.
- Obtain the absorption spectrum of each solution using a laboratory spectrophotometer (e.g., 350 nm to 750 nm, 1 nm increments) as a reference.
3. Optical Analysis Setup:
- Spectrophotometry: Use a commercial instrument for benchmark measurements.
- LED Photometry (PEDD): Construct a system with a specific-wavelength LED source and a matched photodetector LED. The PEDD operates on a charge-discharge cycle principle for high-sensitivity measurement.
- Camera-Based Imaging: Place the sample in a uniform illumination setup (e.g., a lightbox with a diffuser). Use a smartphone mounted on a stand to capture digital images of each sample under identical lighting and camera settings (e.g., fixed focus, white balance, and exposure).
4. Data Processing:
- Spectrophotometry: Record the absorbance at the analyte's peak wavelength.
- LED Photometry: Measure the discharge time of the detector LED, which is inversely proportional to light intensity.
- Camera-Based Imaging: Extract average Red, Green, and Blue (RGB) values from a region of interest (ROI) on the sample image using image analysis software. Convert these values to a quantitative parameter like intensity or Hue.
5. Calibration and Analysis: Plot the measured signal from each method against the reference pH values (or analyte concentration) to generate a calibration curve. Calculate performance metrics like sensitivity, linear dynamic range, and limit of detection from these curves.
The logical workflow for this comparative analysis is outlined below.
Protocol: Single-Molecule Detection with a Smartphone Microscope
This protocol details the methodology for achieving single-molecule fluorescence detection [53].
1. Microscope Assembly:
- Core Components: Construct the microscope using a modular design comprising a protective case, a laser stage (with laser, focusing lens, and alignment screws), an objective stage (with a low numerical aperture air objective and emission filter), and a sample stage (with sample holder and prism).
- Illumination: Employ a laser source (e.g., 640 nm for ATTO 647N) configured for Total Internal Reflection (TIR) or Highly Inclined and Laminated Optical sheet (HILO) illumination to minimize background signal.
- Detection: Use the smartphone's camera, which acts as the tube lens, to focus the emitted light onto its CMOS sensor. An emission filter is crucial to block scattered laser light.
2. Sample Preparation:
- Immobilize fluorescence standards, such as DNA origami structures labeled with a single fluorophore (e.g., ATTO 542 or ATTO 647N), on a clean quartz substrate at a low density suitable for single-molecule imaging.
3. Data Acquisition:
- Place the sample on the microscope stage.
- Using the smartphone's camera in a professional/video mode that allows manual control, record a time-lapse series of images or a video with a suitable exposure time (e.g., 100 ms).
- To observe single-step photobleaching, continuously image the same field of view until the fluorophores bleach.
4. Data Analysis:
- Extract the time-traces of fluorescence intensity from individual spots in the image sequence.
- Identify single molecules by their characteristic single-step photobleaching events.
- Calculate the Signal-to-Noise Ratio (SNR) by dividing the mean signal of a single molecule by the standard deviation of the background.
The Scientist's Toolkit: Essential Research Reagents and Materials
The development and implementation of high-performance smartphone-based optical detectors rely on a set of key materials and reagents. The following table details these essential components and their functions.
Table 2: Key research reagent solutions and materials for smartphone-based optical detection.
| Item | Function/Application | Example Specifications |
|---|---|---|
| Colorimetric pH Dye | Acts as a model analyte for system validation and performance benchmarking. | Bromocresol Green (BCG), 25 µM in solution [75]. |
| Fluorescent Dyes | Label biomolecules for ultrasensitive fluorescence and single-molecule detection. | ATTO 542, ATTO 647N [53]. |
| DNA Origami Structures | Serve as a nanoscale scaffold for precise fluorophore positioning, used as a calibration standard for super-resolution microscopy. | 60 x 52 nm² 2-layer sheet (2LS) with biotins for surface immobilization [53]. |
| Low NA Air Objective | Collects emitted light from the sample in a compact microscope setup. | Inexpensive, finite-conjugation objective [53]. |
| Bandpass Emission Filter | Blocks scattered excitation laser light, allowing only the fluorescence signal to reach the camera. | Filter matched to the fluorophore's emission spectrum [53]. |
| Microfluidic Chip | Provides a platform for automated fluid handling, sample processing, and analysis with small reagent volumes. | Lab-on-a-chip device made from PDMS or similar polymer [19]. |
| 3D-Printed Enclosure | Houses optical components, provides structural stability, and shields the sample from ambient light. | Custom-designed case printed in opaque material [53] [19]. |
Visualization of a Smartphone Microscope for Single-Molecule Detection
The following diagram illustrates the key components and optical path of the smartphone-based microscope capable of single-molecule detection, as described in the experimental protocol [53].
The comparative analysis presented herein clearly indicates that the optimal optical modality for a smartphone-based LOC system is highly application-dependent. For quantitative colorimetric assays where the highest sensitivity and lowest LOD are paramount, the LED Photometry (PEDD) approach demonstrates superior performance [75]. When spatial information or the ubiquity of the detector is the primary concern, camera-based imaging offers a versatile, though less precise, alternative. Remarkably, the convergence of advanced illumination schemes, precise optical design, and the powerful cameras in modern smartphones has enabled single-molecule detection, a capability once confined to research-grade laboratories [53]. As smartphone technology continues to advance, the integration of these optical modalities with microfluidics, smart assays, and artificial intelligence will further democratize powerful molecular analysis, impacting fields from point-of-care diagnostics to environmental monitoring and drug development.
The integration of smartphone-based detection systems with Lab-on-a-Chip (LoC) platforms represents a paradigm shift in point-of-care (PoC) diagnostics, environmental monitoring, and food safety testing [76] [19]. A critical step in translating these novel platforms from research prototypes to trusted analytical tools is their rigorous validation against established gold standard methods, primarily conventional spectrophotometry and laboratory-based Enzyme-Linked Immunosorbent Assay (ELISA) [77]. This process verifies that the portable, often cost-effective smartphone-based systems can deliver performance comparable to that of traditional, expensive laboratory equipment, thereby ensuring data reliability and clinical validity [78] [79]. Framed within the broader context of optical detection methods in smartphone-based LoC research, this technical guide outlines the core principles, methodologies, and analytical frameworks for conducting such validation studies.
Smartphones offer a powerful, integrated package for analytical chemistry, leveraging their high-resolution cameras for optical detection, powerful processors for data analysis, and connectivity for data transmission [19]. These devices are particularly adept at measuring colorimetric and fluorescent signals generated by common bio-assays, such as ELISA, which are traditionally quantified using bulky and expensive microplate readers [76] [77]. The move toward miniaturized, centrifugal microfluidic platforms, like Lab-on-Compact-Disc (LOCD), further underscores the need for compact and portable detection systems that do not compromise on analytical accuracy [77]. This document provides an in-depth examination of the procedures for validating the optical detection components of these emerging systems against their conventional counterparts.
Fundamental Principles of Optical Detection in Smartphone-Based LoC
The core principle behind many smartphone-based optical detectors is absorption spectrophotometry, which is governed by the Beer-Lambert Law [77]. This law establishes a linear relationship between the absorbance (A) of a solution and the concentration of the analyte within it:
A = -log₁₀(I/I₀)
Where:
- A is the absorbance (Optical Density, OD)
- I is the intensity of light transmitted through the sample
- I₀ is the intensity of the incident light [77]
In a conventional microplate reader, a monochromatic light source (e.g., a laser or LED) is passed through the sample, and a dedicated photodetector measures the transmitted light intensity at a specific wavelength, typically 450 nm for assays using the TMB substrate [77]. Smartphone-based systems replicate this function by using their built-in CMOS sensors as the photodetector [80] [19]. Some designs employ an external monochromatic LED to ensure consistent wavelength, while the smartphone camera captures the intensity of light transmitted through the sample chamber on a microfluidic device [77]. The phone's software then converts the captured image data (e.g., RGB values) into an absorbance value based on the Beer-Lambert relationship [80].
Table 1: Comparison of Conventional and Smartphone-Based Optical Detection Platforms
| Feature | Conventional Microplate Reader | Smartphone-Based Reader |
|---|---|---|
| Detection Principle | Absorption spectrophotometry | Absorption spectrophotometry / Image analysis |
| Light Source | Built-in monochromator or LED | External LED or ambient light [77] |
| Detector | Photomultiplier tube or silicon photodiode | CMOS camera sensor [19] |
| Data Processing | Dedicated onboard software | Smartphone application (App) [77] [49] |
| Portability | Low (benchtop instrument) | High (handheld device) [76] |
| Cost | High | Low [19] |
| Throughput | High (96/384 wells) | Variable (depends on LoC design) [81] |
Experimental Design for Validation Studies
A robust validation study must be carefully designed to directly compare the performance of the smartphone-based system with the gold standard method. This involves parallel analysis of the same samples using both platforms.
Sample Preparation and Assay Protocol
For a typical ELISA validation, a set of standards with known analyte concentrations and real-world samples (e.g., patient serum, plasma, or food extracts) are used.
- Standard Curve Generation: A serial dilution of the target analyte (e.g., recombinant protein) is prepared to span the expected dynamic range of the assay. This range should cover from below the expected clinical cutoff to the maximum anticipated concentration [82] [78].
- Parallel Testing: The same set of standards and samples are applied to both the conventional 96-well plate and the LoC platform (e.g., LOCD, paper-based microfluidic chip) [77] [79].
- Assay Execution: The full ELISA protocol—including incubation, washing, and substrate addition—is performed according to the optimized conditions for each platform. For the LOCD, this process is often automated and controlled by centrifugal force [77].
- Signal Measurement: Upon completion, the signal is measured simultaneously or in rapid succession. The gold standard microplate reader measures the absorbance at the requisite wavelength(s). The smartphone-based system captures an image of the LoC device, which is then processed by a dedicated app to determine the absorbance or color intensity for each reaction chamber [80] [77].
Key Validation Parameters
The following core parameters must be evaluated to establish the validity and reliability of the smartphone-LoC system [78] [79]:
- Sensitivity and Limit of Detection (LOD): The lowest concentration of an analyte that the assay can reliably distinguish from zero. It is determined from the standard curve using the formula: LOD = Meanblank + 3SDblank, where SD is the standard deviation of the blank sample [78].
- Precision: The degree of reproducibility of the measurements. This includes:
- Accuracy: The closeness of the measured value to the true value. This is often assessed through spike and recovery experiments, where a known amount of analyte is added to a sample matrix and the measured concentration is compared to the expected value [78] [83]. Recovery rates should ideally be between 80-120%.
- Linearity and Assay Range: The ability of the assay to produce results that are directly proportional to the analyte concentration within a specified range. The standard curve should have a coefficient of determination (R² > 0.98) when fitted with an appropriate model (e.g., 4-parameter logistic, 4PL) [82] [78].
- Specificity: The ability of the assay to detect only the target analyte without cross-reactivity from related substances in the sample matrix [78].
Data Analysis and Correlation Methodologies
Standard Curve Fitting
For quantitative ELISAs, the relationship between absorbance and analyte concentration is often sigmoidal. The 4-parameter logistic (4PL) model is the most widely used and accurate curve fitting method [82]. The model is defined as:
Y = D + (A - D) / (1 + (X/C)^B)
Where:
- Y is the absorbance (OD)
- X is the analyte concentration
- A is the minimum asymptote (background signal)
- D is the maximum asymptote (saturation signal)
- C is the inflection point (EC50)
- B is the slope factor [82]
Both the conventional reader software and the smartphone analysis algorithm should use the same model (4PL) for curve fitting to ensure comparable quantification [82].
Statistical Correlation
The primary statistical method for validation is linear regression analysis between the concentrations measured by the smartphone system (test method) and those measured by the conventional reader (reference method).
- Bland-Altman Plot: This is used to assess the agreement between the two methods by plotting the difference between the measurements against their average. It helps identify any systematic bias or concentration-dependent error [79].
- Passing-Bablok Regression: A non-parametric method that is robust against outliers and non-normal error distribution, making it suitable for comparing clinical measurement methods.
- Calculation of Sensitivity and Specificity: For qualitative or semi-quantitative tests, the diagnostic performance of the smartphone system is evaluated against the gold standard by calculating its clinical sensitivity, specificity, and accuracy [77].
Table 2: Exemplary Validation Data from a Dengue IgG ELISA on an LOCD Platform [77]
| Parameter | Gold Standard Reader | Smartphone-Based LOCD Reader |
|---|---|---|
| Sample Size (n) | 64 | 64 |
| Sensitivity | Benchmark | 95% |
| Specificity | Benchmark | 100% |
| Correlation Coefficient (R²) | 1.00 | >0.98 (vs. Gold Standard) |
| Key Conclusion | - | "High accuracy... when compared with gold standard commercial ELISA microplate readers." |
Case Study: Validation of a Smartphone-Based LOCD ELISA System
A study by Thiha et al. serves as an exemplary model for a thorough validation process [77]. The researchers developed a standalone LOCD platform for performing a dengue antibody IgG ELISA, with results transmitted via Bluetooth to a smartphone.
- Experimental Workflow: The LOCD automated all fluidic steps via centrifugal force. The detection system used a 450 nm LED and a photodiode sensor, with the microcontroller calculating absorbance based on the Beer-Lambert law. Results were displayed on an LCD and sent to a smartphone [77].
- Validation Protocol: The system was tested using 64 patient samples. Each sample was run on the LOCD platform, and the result was compared to that obtained from a commercial ELISA microplate reader.
- Outcome: The smartphone-LOCD system demonstrated 95% sensitivity and 100% specificity, establishing a high level of agreement with the gold standard method [77]. This successful validation underscores the potential of such integrated systems for reliable PoC diagnostics.
The experimental workflow and data correlation process for this validation are summarized in the diagram below.
The Scientist's Toolkit: Essential Research Reagents and Materials
The following reagents, materials, and software are critical for developing and validating smartphone-based LoC systems for optical detection.
Table 3: Essential Research Reagents and Tools for Validation
| Category | Item | Function in Validation |
|---|---|---|
| Assay Components | Coated LoC/Microplate | Solid phase for immobilizing capture antibody/antigen. |
| Matched Antibody Pair | For sandwich ELISA: capture and detection antibodies. | |
| ELISA Standards | Known concentrations of analyte for generating the standard curve. | |
| TMB Substrate | Enzyme substrate that produces a colorimetric product measurable at 450 nm. | |
| Stop Solution (e.g., H₂SO₄) | Halts the enzymatic reaction and stabilizes the final color. | |
| Sample & Buffer | Biological Samples | Serum, plasma, etc., for testing in a real-world matrix. |
| Assay Buffer / Diluent | Matrix for reconstituting standards and diluting samples. | |
| Blocking Buffer (e.g., BSA) | Prevents non-specific binding to the solid surface. | |
| Hardware | Smartphone | Serves as detector, processor, and data interface. |
| Microplate Reader | Gold standard instrument for absorbance measurement. | |
| Monochromatic LED (e.g., 450 nm) | Provides consistent light source for smartphone detection [77]. | |
| Software & Analysis | 4PL/5PL Curve Fitting Software | For accurate interpolation of sample concentrations. |
| Statistical Software (e.g., R, GraphPad Prism) | To perform regression and correlation analyses. | |
| Custom Smartphone App | Controls detection, processes images, and calculates results. |
Common Pitfalls and Troubleshooting in Validation
Several challenges can arise during the validation of smartphone-based systems:
- Matrix Effects: Complex sample matrices (e.g., blood, food homogenates) can cause interference, leading to inaccurate readings. Solution: Perform spike and recovery experiments and use matrix-matched standards for the calibration curve to correct for these effects [78] [83].
- High Background Signal: This can lead to false positives or reduced sensitivity. Solution: Optimize blocking buffers and washing protocols to minimize non-specific binding [78].
- Poor Reproducibility Between Smartphones: Different smartphone models have varying camera sensors and image processing algorithms. Solution: Use an external, calibrated light source and develop device-agnostic analysis algorithms that rely on internal controls and calibration within each image [80] [81].
- Inadequate Curve Fitting: Using a linear model for a non-linear (sigmoidal) ELISA response will yield inaccurate results. Solution: Always use a 4- or 5-parameter logistic (4PL/5PL) regression model for fitting the standard curve [82].
The validation of smartphone-based LoC systems against gold standard spectrophotometry and ELISA is not merely a procedural step but a fundamental requirement for ensuring data integrity and fostering adoption in research, clinical, and field settings. By adhering to rigorous experimental design, employing appropriate statistical correlation methods, and systematically addressing common pitfalls, researchers can confidently demonstrate that these innovative, portable platforms are capable of performance on par with traditional laboratory equipment. As smartphone technology and microfluidic design continue to advance, the principles of validation outlined in this guide will remain central to the development of robust, reliable, and accessible analytical tools for the future.
The integration of smartphones as the core analytical platform in lab-on-a-chip (LOC) systems represents a paradigm shift in portable molecular diagnostics. By leveraging the integrated cameras and computational power of smartphones, these systems offer a viable path toward democratizing chemical and biological analysis for point-of-care, environmental monitoring, and food safety applications. This whitepaper provides a critical evaluation of the real-world performance of smartphone-based LOC devices, with a specific focus on user-friendliness, analytical throughput, and application scope. Framed within the broader principles of optical detection methods, this guide details experimental methodologies for quantifying performance metrics, supported by structured data and standardized protocols. The convergence of microfluidics, nanomaterials, and smartphone technology holds immense promise for creating powerful, versatile, and accessible analytical tools that are no longer confined to the laboratory.
The push to translate laboratory-grade analyses from centralized facilities to the point-of-need is a major driver in LOC research. Traditional microfluidic systems often require bulky, expensive peripheral equipment for operation and detection, negating the benefits of miniaturization. Smartphones present a transformative solution by integrating a powerful suite of optoelectronic components, processors, and connectivity features into a single, globally ubiquitous device [19]. Their cameras are sophisticated optical sensors capable of various detection modalities, including fluorescence, colorimetry, and surface-enhanced Raman scattering (SERS). Furthermore, the integrated flash serves as a high-intensity light source, and onboard computational resources can run custom analysis applications and even machine learning algorithms for data interpretation [19] [47].
The motivations for adopting smartphones are multifaceted. With over 4.69 billion users globally and a market that drives rapid technological advancement and cost reduction, smartphones offer an economy of scale unattainable by bespoke scientific instruments [19] [84]. This review establishes a framework for evaluating these systems based on three critical pillars that determine their translational potential: User-Friendliness (the ease of operation by non-experts), Throughput (the number of analyses per unit time), and Application Scope (the range of analytes and settings in which the device is effective).
Principles of Optical Detection in Smartphone-Based LOC
The primary interface between the smartphone and the biochemical assay in an LOC device is the camera, which is used to detect optical signals generated or modulated by the presence of a target analyte.
- Colorimetry and Absorbance: These methods measure the color intensity or light absorption of a sample, often linked to enzyme-linked immunosorbent assays (ELISA) or plasmonic changes in gold nanoparticles. The smartphone camera captures images of the detection zone, and the intensity of the red, green, and blue (RGB) color channels is quantified to determine analyte concentration [19] [47].
- Fluorescence: This method relies on light emission from fluorophores at a specific wavelength after excitation. Smartphone-based fluorescence detection often requires external filters to block the excitation light. Signal enhancement techniques, such as Metal-Enhanced Fluorescence (MEF) using gold or silver nanoparticles, are frequently employed to boost sensitivity to detectable levels for smartphone cameras [47].
- Surface-Enhanced Raman Scattering (SERS): SERS provides a unique molecular fingerprint by dramatically enhancing the Raman signal of molecules adsorbed onto nano-structured metal surfaces. Smartphones can be adapted to read SERS spectra using simplified spectrometers, enabling highly specific and sensitive multiplexed detection [47] [85].
The following workflow diagram illustrates the general process of optical detection and analysis using a smartphone-based LOC system.
Optical Detection Workflow in Smartphone-Based LOC
Evaluating User-Friendliness
User-friendliness is paramount for the deployment of technology in resource-limited or non-laboratory settings. It encompasses the simplicity of the hardware setup, the intuitiveness of the software, and the minimal requirement for user intervention.
Key Metrics and Evaluation Methods
Table 1: Metrics for Evaluating User-Friendliness
| Metric | Description | Evaluation Method |
|---|---|---|
| Assay Steps | Number of manual preparatory and operational steps before reading. | Protocol analysis; count of user-dependent fluidic handling steps (e.g., pipetting, mixing, incubation) [86]. |
| Setup Time | Time required from unboxing the device to obtaining a result. | Timed trials with naive users; average the results from multiple trials [19]. |
| Intuitiveness of App UI | Clarity of the application user interface and result reporting. | Heuristic evaluation by UX experts; user surveys with Likert scales rating ease of navigation and clarity of instructions [19]. |
| Hardware Integration | Complexity of assembling the smartphone with the LOC accessory. | Binary assessment (Integrated/Modular); count of mechanical attachments and optical alignments required [19] [87]. |
Experimental Protocol for Usability Testing
- Recruitment: Recruit a cohort of n ≥ 15 users representing the target audience (e.g., field technicians, nurses, community workers). Include individuals with no prior experience in analytical chemistry.
- Training: Provide a standardized, brief (e.g., 5-minute) overview of the device's purpose. Do not provide detailed, step-by-step instructions for operation.
- Task Execution: Ask each user to perform a complete analysis of a standardized sample using the smartphone-LOC system. Record the time taken from start to finish.
- Data Collection: Have users complete a post-test questionnaire rating the following on a scale of 1 (Very Difficult) to 5 (Very Easy):
- Ease of physical setup.
- Clarity of the smartphone application instructions.
- Confidence in interpreting the final result.
- Data Analysis: Calculate the average task completion time and the average score for each survey question. A successful device should achieve an average score of ≥4.0 on usability questions.
Evaluating Throughput
Throughput determines the capacity of a system to handle multiple samples or analytes, which is crucial for widespread screening and monitoring applications.
Key Metrics and Evaluation Methods
Table 2: Metrics for Evaluating Throughput
| Metric | Description | Evaluation Method |
|---|---|---|
| Analysis Time | Time from sample introduction to result output, including any incubation or processing. | Direct measurement with a stopwatch during controlled experiments; report mean ± standard deviation from n=5 replicates. |
| Samples per Hour | The total number of individual samples that can be processed in one hour. | Calculated as (60 minutes / Total Analysis Time per Sample). For parallel processing, this value is multiplied by the number of parallel channels [19] [86]. |
| Multiplexing Capacity | The number of distinct analytes that can be detected simultaneously from a single sample. | Determined by the assay design (e.g., number of different capture zones on a lateral flow strip, spectral distinctness of SERS tags, or number of fluorescence channels) [47]. |
Experimental Protocol for Throughput Assessment
- Baseline Timing: Using a pre-prepared and optimized system, load a single sample and start the timer. Record the time until the result is displayed on the smartphone screen. Repeat this process five times to establish a baseline analysis time.
- Parallel Processing Evaluation: If the LOC device is designed to process multiple samples in parallel (e.g., a multi-channel microfluidic chip), repeat Step 1 but by loading all channels simultaneously. The timer stops when results for all channels are available.
- Throughput Calculation:
- For serial devices: Throughput (samples/hour) = 3600 / Average Analysis Time (s).
- For parallel devices: Throughput (samples/hour) = [Number of Parallel Channels] * [3600 / Total Analysis Time for the full batch (s)].
- Multiplexing Verification: Prepare a sample containing a known concentration of three different target analytes. Run the assay and verify that the smartphone application can independently and correctly identify and quantify all three targets without cross-talk.
Evaluating Application Scope
The application scope defines the technological boundaries and practical utility of a smartphone-LOC system across different fields and sample matrices.
Key Domains and Enabling Technologies
Table 3: Diverse Application Scopes of Smartphone-Based LOC Systems
| Application Domain | Target Analytes | Common Optical Detection Method | Key Enabling Technology |
|---|---|---|---|
| Clinical Diagnostics | Disease biomarkers (proteins, nucleic acids), viruses | Fluorescence, Colorimetric (LFA) | Nanoparticles for signal enhancement (MEF, SERS); microfluidics for sample preparation [19] [47]. |
| Food Safety & Agriculture | Pathogens (E. coli, Salmonella), pesticides, toxins | Colorimetry, Fluorescence | Nanomaterial-based LOCs (e.g., graphene electrodes); immunoassays integrated into microfluidic chips [86]. |
| Environmental Monitoring | Heavy metal ions, organic pollutants, water pH | Colorimetry, SERS | Plasmonic nanoparticles (for SERS); specific colorimetric probes [47] [86]. |
| Infrastructure Monitoring | 3D structural data | LiDAR, RGB Photogrammetry | Integrated LiDAR sensors; RTK rover accessories for geolocation [87]. |
Experimental Protocol for Assessing Analytical Figures of Merit
To standardize the reporting of performance across different applications, the following protocol should be used to characterize a smartphone-LOC system's core analytical capabilities.
Limit of Detection (LOD) Determination:
- Prepare a dilution series of the target analyte in the relevant sample matrix (e.g., serum, river water, food homogenate) spanning concentrations from blank to well above the expected detection limit.
- Analyze each concentration with n=5 replicates using the smartphone-LOC system.
- Plot the measured signal (e.g., RGB intensity, pixel count) against the analyte concentration.
- Calculate the LOD using the formula: LOD = (3.3 × σ) / S, where σ is the standard deviation of the blank sample's signal and S is the slope of the calibration curve.
Dynamic Range and Sensitivity:
- From the calibration curve generated in Step 1, the dynamic range is defined as the concentration interval over which the signal has a linear relationship (R² > 0.99) with the analyte concentration. Sensitivity is the slope of this linear region.
Specificity/Selectivity Assessment:
- Test the system with samples containing potential interfering substances that are likely to be present in the real sample matrix.
- The signal generated by a sample containing the target analyte should be significantly greater (e.g., >90% signal inhibition in competitive assays) than the signal from samples containing only interferents at their expected maximum concentration.
The following diagram outlines the logical decision process for selecting an appropriate optical detection method based on the requirements of the target application.
Optical Detection Method Selection Logic
The Scientist's Toolkit: Essential Research Reagents and Materials
The development and operation of high-performance smartphone-LOC devices rely on a suite of specialized reagents and materials.
Table 4: Key Research Reagent Solutions for Smartphone-Based LOC
| Item | Function | Example in Context |
|---|---|---|
| Gold Nanoparticles (AuNPs) | Plasmonic transducers for colorimetric assays and MEF; provide a surface for bio-conjugation. | Spherical AuNPs for LFA; gold nanostars to create electromagnetic "hotspots" for enhanced fluorescence or SERS [47]. |
| Silver Nanoparticles (AgNPs) | Generate strong localized surface plasmon resonance (LSPR) fields for high-efficiency MEF. | Used in fluorescence microarrays for viral DNA detection, achieving significant signal amplification [47]. |
| Fluorescent Dyes & Quantum Dots | Generate signal for fluorescence-based detection; offer tunable emission wavelengths. | Coupled with antibodies or DNA probes for specific target recognition; used in multiplexing via different emission colors [47]. |
| Microfluidic Chip Substrates | Form the physical structure of the LOC, containing microchannels and reaction chambers. | Polymers (e.g., PDMS, PMMA) for low-cost fabrication; glass for high chemical stability and optical clarity [86]. |
| Surface Modification Reagents | Functionalize surfaces to immobilize capture probes (antibodies, aptamers) and reduce non-specific binding. | Silane chemistry for glass/oxide surfaces; thiolated DNA or PEG molecules for gold surfaces [47]. |
| SERS Tags | Provide intense, characteristic Raman signals for highly specific and multiplexed detection. | Noble metal nanoparticles coated with a Raman reporter molecule and a protective layer [47]. |
A rigorous and standardized approach to evaluating user-friendliness, throughput, and application scope is critical for advancing smartphone-based LOC technologies from compelling research prototypes to robust tools for real-world analysis. The experimental frameworks and metrics outlined in this whitepaper provide a foundation for cross-platform comparison and meaningful performance validation. The ongoing convergence of smartphones with advanced nanomaterials, sophisticated microfluidics, and intelligent software promises a future for molecular analysis that is not only powerful and versatile but also truly democratized and accessible beyond traditional laboratory settings. Future research must focus on overcoming the remaining challenges in sample preparation automation, reagent stability, and large-scale manufacturing to fully realize this potential.
The integration of optical detection methods into smartphone-based lab-on-a-chip (LoC) systems presents researchers with a complex landscape of analytical techniques. This whitepaper establishes a structured decision framework to guide the selection of the most appropriate method based on specific analytical requirements, sample properties, and operational constraints. By synthesizing current advancements in microfluidic technologies and biosensing platforms, we provide a systematic approach to method optimization that balances sensitivity, cost, portability, and technical feasibility for drug development and scientific research applications. The framework specifically addresses the unique opportunities and challenges presented by smartphone-integrated systems, which provide computational power, wireless connectivity, and high-resolution imaging capabilities to LoC platforms [60].
The rise of smartphone-based analytical devices represents a paradigm shift in point-of-care (PoC) in vitro medical diagnostics and environmental monitoring. These systems combine the miniaturization and efficiency of LoC technologies with the ubiquitous processing power and connectivity of consumer mobile devices [88]. Within this context, optical detection methods have emerged as particularly versatile tools for researchers developing next-generation analytical platforms.
Despite their growing adoption, researchers face significant challenges in selecting the optimal optical method for specific applications. The decision process involves evaluating multiple interdependent variables including target analytes, matrix effects, sensitivity requirements, and resource constraints. This complexity is further compounded by the rapid evolution of nanomaterials, imaging technologies, and data analytics algorithms [60]. Without a structured selection framework, researchers risk suboptimal system performance, increased development costs, and delayed project timelines.
This technical guide addresses this critical gap by presenting a comprehensive decision framework specifically tailored for smartphone-integrated LoC systems. By combining comparative analysis of technical parameters with structured workflow visualizations, we empower researchers to make informed, systematic decisions that align method selection with specific analytical needs and operational contexts.
Comparative Analysis of Optical Detection Methods
Selecting an appropriate detection method requires a clear understanding of the technical capabilities and limitations of available technologies. The following analysis focuses on methods most compatible with smartphone-based LoC platforms, emphasizing their suitability for resource-constrained or field-based settings.
Technical Performance Metrics
Table 1: Performance Characteristics of Primary Optical Detection Methods
| Detection Method | Typical LOD | Dynamic Range | Analysis Time | Multiplexing Capability | Cost Index |
|---|---|---|---|---|---|
| Absorbance | μM-nM | 3-4 orders | Minutes | Low | Low |
| Fluorescence | pM-fM | 4-6 orders | Minutes-Hours | Medium-High | Medium |
| Chemiluminescence | fM-aM | 4-5 orders | Seconds-Minutes | Medium | Low-Medium |
| Surface Plasmon Resonance | nM-pM | 3-4 orders | Minutes | Low | High |
The data reveals a clear trade-off between sensitivity and cost/complexity. Fluorescence-based methods offer exceptional sensitivity (pM-fM) and are widely employed in smartphone-based detection due to the high-quality cameras available on modern devices [60]. Chemiluminescence provides superior sensitivity without requiring an excitation light source, significantly simplifying optical design [60]. Absorbance spectroscopy, while less sensitive, remains valuable for applications involving higher analyte concentrations and offers advantages in cost and simplicity.
Smartphone Integration Considerations
Table 2: Smartphone Integration Requirements and Complexities
| Method | Optical Components Required | Data Processing Demand | Power Requirements | Ease of Miniaturization |
|---|---|---|---|---|
| Absorbance | LED, lens, filter | Low | Low | High |
| Fluorescence | LED/laser, emission filter, lens | Medium-High | Medium-High | Medium |
| Chemiluminescence | Lens (no light source) | Low | Low | High |
| SPR | Polarizer, lens, specialized chip | High | Medium | Low |
When integrated with smartphones, each method presents distinct engineering challenges. Fluorescence detection requires careful optical alignment and filtering to separate excitation and emission signals, but leverages the smartphone's sophisticated imaging capabilities [60]. Chemiluminescence and absorbance measurements benefit from simpler optical arrangements, making them particularly suitable for field-deployable devices where robustness and simplicity are prioritized [60]. Surface Plasmon Resonance (SPR) offers label-free detection but typically requires more sophisticated optical components that challenge miniaturization efforts.
Decision Framework Methodology
The selection of an optimal optical detection method requires systematic evaluation of multiple technical and operational factors. The following structured approach guides researchers through this decision process.
Primary Decision Algorithm
The foundational decision path begins with assessing core analytical requirements, then progressively incorporates constraints related to sample properties and operational environment.
Secondary Optimization Pathway
After identifying a primary method through the initial algorithm, this secondary pathway addresses key implementation considerations to optimize performance and practicality.
Experimental Protocol Specifications
Implementing the selected method requires careful attention to protocol details. The following specifications correspond to key decision points in the framework.
Fluorescence-Based Detection Protocol
For applications requiring high sensitivity and multiplexing capability (following Path 1 in the Decision Algorithm):
Sample Preparation:
- Dilute sample in appropriate buffer (e.g., PBS, pH 7.4)
- Add fluorescence probe (antibody, aptamer, or molecular beacon) at optimized concentration
- Incubate for 15-30 minutes at room temperature
Nanomaterial Enhancement:
- Incorporate gold nanoparticles (AuNPs) or quantum dots to enhance signal intensity [60]
- Functionalize nanomaterials with appropriate recognition elements (antibodies, aptamers)
- Optimize nanoparticle concentration to maximize signal-to-noise ratio
Smartphone Imaging:
- Utilize built-in flash as excitation source
- Implement external bandpass filters to separate excitation and emission wavelengths
- Capture images using manual mode with fixed ISO, exposure, and white balance settings
- Process images using dedicated mobile application for intensity quantification
Absorbance-Based Detection Protocol
For applications where moderate sensitivity is sufficient and cost simplicity is prioritized (following Path 2 in the Decision Algorithm):
Sample Preparation:
- Mix sample with colorimetric reagent (enzyme substrate, pH indicator, or specific chromogen)
- Incubate for 5-10 minutes for color development
Microfluidic Integration:
- Design channel depth for adequate path length (typically 0.5-1.0 mm)
- Incorporate mixing structures for efficient reagent-sample interaction
Smartphone Detection:
- Utilize white LED source for uniform illumination
- Capture transmitted light intensity through sample
- Convert RGB values to absorbance using logarithmic transformation
- Apply color space transformation (RGB to HSV) for more robust quantification
Essential Research Reagent Solutions
The successful implementation of optical detection methods in smartphone-based LoC platforms relies on carefully selected reagents and materials. The following table summarizes key components and their functions in developing these analytical systems.
Table 3: Key Research Reagents and Materials for Smartphone-Based LoC Development
| Reagent/Material | Function | Application Examples | Considerations |
|---|---|---|---|
| Gold Nanoparticles (AuNPs) | Signal amplification, quenching agent, plasmonic enhancement | Fluorescence enhancement, colorimetric assays, SPR | Tunable optical properties, high surface-to-volume ratio [60] |
| Graphene Oxide (GO) | Quenching platform, adsorption substrate, electrode material | Fluorescence quenching assays, electrochemical detection | Large surface area with oxygen functional groups [60] |
| Quantum Dots | Fluorescent labels with high quantum yield | Multiplexed detection, tracking | Narrow emission peaks, photostability |
| Molecularly Imprinted Polymers (MIPs) | Synthetic recognition elements | Chemical contaminant detection, small molecule sensing | High stability, customizable binding sites [60] |
| Aptamers | Recognition elements | Pathogen detection, protein biomarkers | Thermal stability, chemical modification capability [60] |
| Enzymes (HRP, ALP) | Signal generation through substrate conversion | Chemiluminescence, colorimetric assays | High catalytic efficiency, substrate specificity [60] |
Advanced Implementation Considerations
Beyond the initial selection of detection methodology, several advanced factors critically influence the performance and reliability of smartphone-based LoC systems.
Microfluidic Integration
The interface between optical detection and microfluidic delivery systems requires careful engineering to maximize analytical performance:
- Optical Path Length: For absorbance measurements, channel depth must be optimized to balance sensitivity against sample volume requirements and illumination uniformity [60].
- Surface Interactions: Nonspecific adsorption to channel walls can interfere with both fluorescence and label-free detection methods, requiring appropriate surface treatments or blocking agents.
- Flow Control: Precisely controlled flow rates are essential for consistent incubation times, thorough mixing, and reproducible measurements across samples.
Data Processing and Analytics
Smartphone integration enables sophisticated data processing approaches that can enhance analytical performance:
- Image Analysis Algorithms: Implement background subtraction, flat-field correction, and intensity normalization to compensate for environmental variability [60].
- Machine Learning Integration: Utilize smartphone computational resources for pattern recognition in multiplexed assays and classification of complex sample types [60].
- Cloud Connectivity: Transfer raw data to cloud resources for more intensive processing while maintaining a lightweight mobile application [88].
System Evolvability and Platform-Based Design
Adopting a platform-based design (PBD) methodology enhances the long-term viability of smartphone-LoC systems by supporting managed transitions between system generations [88]. Key strategies include:
- Modular Hardware Accessories: Design interchangeable components that interface with a common smartphone platform to support multiple detection modalities [88].
- Abstracted Software Layers: Implement change-absorbing interfaces in software architecture to accommodate new LoC variants without fundamental redesign [88].
- Standardized Communication Protocols: Establish consistent data exchange formats between smartphone, hardware accessory, and LoC components to support future expansion.
This decision framework provides a systematic methodology for selecting optimal optical detection methods in smartphone-based LoC research. By integrating technical performance metrics with practical implementation constraints, the framework guides researchers through a structured evaluation process that balances analytical sensitivity, operational requirements, and resource limitations. The integration of standardized workflow visualizations, performance comparison tables, and detailed reagent specifications offers a comprehensive reference tool for scientists developing next-generation analytical platforms. As smartphone-LoC technologies continue to evolve, this structured selection approach will enable more efficient development of robust, high-performance systems for pharmaceutical research, clinical diagnostics, and environmental monitoring applications.
Conclusion
Smartphone-based optical detection for Lab-on-a-Chip systems represents a paradigm shift towards decentralized, accessible, and powerful molecular analysis. By synthesizing key insights, this review establishes that the convergence of smartphone technology with advanced optical methods and AI-powered data analysis can achieve remarkable sensitivity, in some cases down to the single-molecule level. Future progress hinges on overcoming persistent challenges in standardization, system integration, and manufacturing scalability. The ongoing innovation in this field promises to profoundly impact biomedical research and clinical practice, enabling new frontiers in personalized medicine, rapid diagnostics, and real-time health monitoring in both resource-rich and limited settings.
References
- 1. Optical biosensors - PMC [https://pmc.ncbi.nlm.nih.gov/articles/PMC4986466/]
- 2. Smartphone-Based Biosensors: Current Trends ... [https://www.mdpi.com/2673-4591/106/1/10]
- 3. OPTICAL DETECTION TECHNIQUES FOR MICROFLUIDICS [https://elveflow.com/microfluidic-reviews/optical-detection-techniques-microfluidics/]
- 4. Colorimetric Detection - an overview [https://www.sciencedirect.com/topics/physics-and-astronomy/colorimetric-detection]
- 5. Detection Methods - Yale Center for Molecular Discovery | [https://ycmd.yale.edu/instrumentation-and-libraries/plate-readers/detection-methods]
- 6. A smartphone label-free and automated thermo-analytical ... [https://www.sciencedirect.com/science/article/abs/pii/S004896972408272X]
- 7. Label-free biodetection using a smartphone [https://pubmed.ncbi.nlm.nih.gov/23609514/]
- 8. Enhanced optical imaging and fluorescent labeling for ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC11143489/]
- 9. Recent Advances in Smart Phone-Based Biosensors for Various Applications [https://www.mdpi.com/2227-9040/13/7/221]
- 10. Smartphone biosensors - Nanosensors Group [https://nano.ece.illinois.edu/research-smartphone-biosensors/smartphone-biosensors/]
- 11. Applications of smartphone-based colorimetric biosensors [https://www.sciencedirect.com/science/article/pii/S2590137022000681]
- 12. Smartphone-based mobile biosensors for the point-of-care testing of human metabolites - PMC [https://pmc.ncbi.nlm.nih.gov/articles/PMC9034388/]
- 13. Cradle of Optical Components Turns Smartphone into ... [https://www.techbriefs.com/component/content/article/30628-cradle-of-optical-components-tu]
- 14. Smartphone-Based Multiplexed Biosensing Tools for Health Monitoring - PMC [https://pmc.ncbi.nlm.nih.gov/articles/PMC9406027/]
- 15. Smartphone-Based Biosensor Devices for Healthcare [https://pmc.ncbi.nlm.nih.gov/articles/PMC8945789/]
- 16. Wireless electrochemiluminescent biosensors: Powering ... [https://pubmed.ncbi.nlm.nih.gov/38387231/]
- 17. Portable on-chip colorimetric biosensing platform ... [https://www.nature.com/articles/s41598-021-02580-w]
- 18. Smartphone-based platforms implementing microfluidic ... [https://www.nature.com/articles/s41467-023-36017-x]
- 19. Smartphones as a platform for molecular analysis [https://pubs.rsc.org/en/content/articlehtml/2025/lc/d4lc00966e]
- 20. Microfluidic Sensors Integrated with Smartphones for ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12298308/]
- 21. Portable 3D-Printed Paper Microfluidic System with a ... [https://www.mdpi.com/2227-9040/13/2/51]
- 22. 3D Printed Auto-mixing Chip Enables Rapid Smartphone ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC6347025/]
- 23. Smartphone-based mobile biosensors for the point-of-care ... [https://www.sciencedirect.com/science/article/pii/S2590006422000527]
- 24. MEMS technology in analytical - Analyst (RSC Publishing)... chemistry [https://pubs.rsc.org/en/content/articlehtml/2003/an/b211229a]
- 25. Effectiveness of smartphone technology for detection ... [https://pubmed.ncbi.nlm.nih.gov/40448047/]
- 26. Cost-Effectiveness Analysis in Implementation Science [https://pmc.ncbi.nlm.nih.gov/articles/PMC7980756/]
- 27. Green analytical chemistry: integrating sustainability into ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC11772533/]
- 28. Green analytical chemistry approaches on environmental ... [https://www.sciencedirect.com/science/article/abs/pii/S2214158822000046]
- 29. Green Analytical Chemistry—Recent Innovations [https://www.mdpi.com/2673-4532/6/1/10]
- 30. Smartphone-Enabled Colorimetry - PMC [https://pmc.ncbi.nlm.nih.gov/articles/PMC10304433/]
- 31. Smartphone based colorimetric approach for quantitative ... [https://www.nature.com/articles/s41598-023-48962-0]
- 32. Portable and Rapid Smartphone - Based of... Colorimetric Assay [https://pubmed.ncbi.nlm.nih.gov/40649312/]
- 33. Fluorescence Fundamentals | Thermo Fisher Scientific - US [https://www.thermofisher.com/us/en/home/references/molecular-probes-the-handbook/introduction-to-fluorescence-techniques.html]
- 34. Single-molecule fluorescence microscopy review [https://pmc.ncbi.nlm.nih.gov/articles/PMC5520217/]
- 35. Review Highly sensitive fluorescence detection technology ... [https://www.sciencedirect.com/science/article/abs/pii/S1359644603027521]
- 36. Comparison of enzyme-linked immunosorbent assay with ... [https://pubmed.ncbi.nlm.nih.gov/2981902/]
- 37. A performance comparison between fluorescent ... [https://www.nature.com/articles/s41598-022-21581-x]
- 38. What Is Single Molecule Microscopy? [https://www.teledynevisionsolutions.com/learn/learning-center/scientific-imaging/what-is-single-molecule-microscopy/]
- 39. The Fluorescent Protein Color Palette [https://evidentscientific.com/en/microscope-resource/knowledge-hub/techniques/confocal/applications/fpcolorpalette]
- 40. The fluorescent protein color palette [https://pubmed.ncbi.nlm.nih.gov/18228502/]
- 41. Highly sensitive fluorescence detection technology ... [https://pubmed.ncbi.nlm.nih.gov/12867149/]
- 42. Label‐free detection techniques for protein microarrays [https://pmc.ncbi.nlm.nih.gov/articles/PMC7167936/]
- 43. Label and Label-Free Detection Techniques for Protein ... [https://www.mdpi.com/2076-3905/4/2/228]
- 44. Label-free single-molecule optical detection | npj Biosensing [https://www.nature.com/articles/s44328-025-00048-9]
- 45. Label-free interaction analysis [https://www.malvernpanalytical.com/en/products/measurement-type/label-free-analysis]
- 46. Label-free optical biosensors in the pandemic era - PMC [https://pmc.ncbi.nlm.nih.gov/articles/PMC11502114/]
- 47. Multiplexed Optical Nanobiosensing Technologies for ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12562344/]
- 48. Label -free measuring biomolecular interactions using ... [https://www.sciencedirect.com/science/article/abs/pii/S0925400524009602]
- 49. Smartphone-Based Multiplexed Biosensing Tools for ... [https://www.mdpi.com/2079-6374/12/8/583]
- 50. Smartphones as Optical Detectors in Pharmaceutical ... [https://www.sciencedirect.com/science/article/abs/pii/S0026265X2500671X]
- 51. Smartphones as a platform for molecular analysis [https://pubs.rsc.org/en/content/articlelanding/2025/lc/d4lc00966e]
- 52. Super-resolution microscopy [https://en.wikipedia.org/wiki/Super-resolution_microscopy]
- 53. Direct single-molecule detection and super-resolution ... [https://www.nature.com/articles/s41467-025-63993-z]
- 54. 5 advantages of digital assays that will dramatically ... [https://idmxs.org/blog/5-advantages-of-digital-assays-that-will-dramatically-accelerate-analytics-and-diagnostics/]
- 55. Point-of-Care Detection of Carcinoembryonic Antigen (CEA ... [https://pubmed.ncbi.nlm.nih.gov/39727865/]
- 56. A smartphone-based fiber-optic aptasensor for label-free ... [https://pubs.rsc.org/en/content/articlelanding/2020/ay/c9ay02406a]
- 57. Direct Single-Molecule Detection and Super Resolution ... [https://moorfield.co.uk/applications/multilayer-composite/]
- 58. A Portable Optical Sensor for Microplastic Detection [https://www.mdpi.com/2076-3417/15/9/4757]
- 59. An innovative eco-friendly optical sensor designed ... [https://www.sciencedirect.com/science/article/pii/S2214180424000692]
- 60. Smartphone-Integrated Electrochemical Devices for ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12467198/]
- 61. Environmental Monitoring: A Comprehensive Review on ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC9688474/]
- 62. A quantitative framework for the analysis of multimodal ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC5337182/]
- 63. Controlled Lighting and Illumination-Independent Target ... [https://www.mdpi.com/1424-8220/19/6/1390]
- 64. Machine vision with flickerless LED lighting [https://antmicro.com/blog/2022/02/machine-vision-with-flickerless-led-lighting/]
- 65. Intelligent Luminance Control of Lighting Systems Based ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC5336081/]
- 66. Automated smartphone based cell analysis platform [https://www.nature.com/articles/s44303-025-00093-z]
- 67. Smartphone-Assisted Thin-Layer Chromatography for Rapid ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12197440/]
- 68. Nanobiosensors for Single-Molecule Diagnostics [https://pmc.ncbi.nlm.nih.gov/articles/PMC12564644/]
- 69. Multiplexed Optical Nanobiosensing Technologies for ... [https://www.mdpi.com/2079-6374/15/10/682]
- 70. Nanobiosensors for Single-Molecule Diagnostics: Toward ... [https://www.preprints.org/manuscript/202509.1525/v1]
- 71. Emerging trends in AI-integrated optical biosensors for point ... [https://pubs.rsc.org/en/content/articlehtml/2025/cc/d5cc04899k]
- 72. IoT powered RNN for improved human activity recognition ... [https://www.nature.com/articles/s41598-025-94689-5]
- 73. A Review of Indoor Localization Methods Leveraging ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC11548474/]
- 74. Optical design is integral in today's medical products [https://flex.com/resources/eye-on-optics-optical-design-is-integral-in-todays-medical-products/]
- 75. A Comparative Study of Optical Sensing Methods for ... [https://www.mdpi.com/1424-8220/25/13/3850]
- 76. Novel Developments of Mobile Sensing Based on the ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC5142836/]
- 77. A Colorimetric Enzyme-Linked Immunosorbent Assay ... [https://www.mdpi.com/1424-8220/15/5/11431]
- 78. ELISA Assay Validation: Best Practices, Methods, and Key ... [https://www.nebiolab.com/elisa-assay-validation-guide/]
- 79. Validation of the ELISA Method for Quantitative Detection of ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC5661705/]
- 80. Smartphone-based optical assays in the food safety field [https://www.sciencedirect.com/science/article/pii/S0165993620301631]
- 81. Smartphone-based optical assays in the food safety field [https://pmc.ncbi.nlm.nih.gov/articles/PMC7457721/]
- 82. How to Analyze ELISA Data and Calculate Results [https://www.bosterbio.com/blog/post/how-to-analyze-elisa-data-and-calculate-results-step-by-step-guide-with-troubleshooting-tips?srsltid=AfmBOoqRG3ZGkdmgyOFYjpKFQjPUbTJAFikvjgKKyvER9HHr-k5C86dP]
- 83. ELISA Guide; Part 3: ELISA Optimization [https://www.jacksonimmuno.com/secondary-antibody-resource/immuno-techniques/elisa-guide-part-3-elisa-optimization/]
- 84. Smartphone Usage Statistics for 2025 (Surprising) [https://backlinko.com/smartphone-usage-statistics]
- 85. Next-generation photonic sensors for single-molecule ... [https://www.sciencedirect.com/science/article/pii/S2709472325000486]
- 86. Recent developments and applications of nanomaterial ... [https://www.sciencedirect.com/science/article/abs/pii/S0924224423001206]
- 87. Evaluation of Positioning Accuracy Using Smartphone RGB ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12252411/]
- 88. Evolvable Smartphone-Based Platforms for Point-of-Care ... [https://www.mdpi.com/2075-4418/6/3/33]